Validating LSER Predictions for LDPE/Water Partitioning: A Robust Framework for Pharmaceutical Safety and Risk Assessment
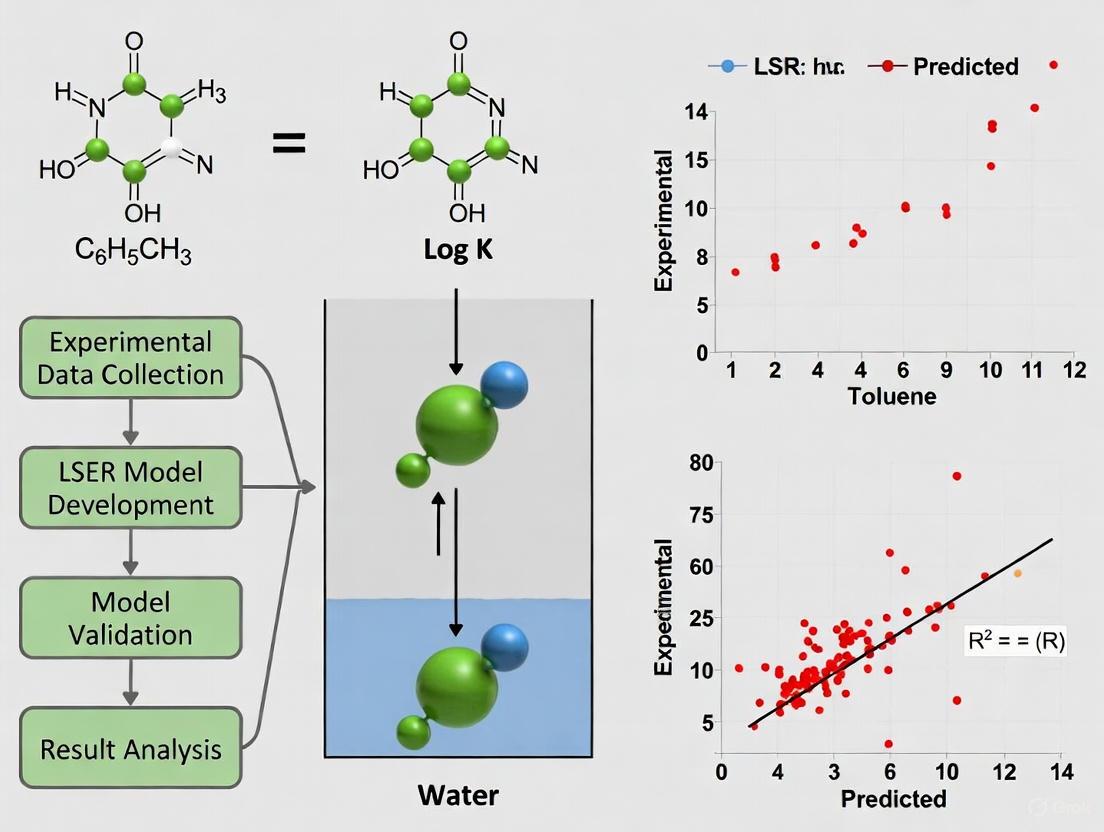
This article provides a comprehensive overview of the validation and application of Linear Solvation Energy Relationship (LSER) models for predicting partition coefficients between low-density polyethylene (LDPE) and water.
Validating LSER Predictions for LDPE/Water Partitioning: A Robust Framework for Pharmaceutical Safety and Risk Assessment
Abstract
This article provides a comprehensive overview of the validation and application of Linear Solvation Energy Relationship (LSER) models for predicting partition coefficients between low-density polyethylene (LDPE) and water. Tailored for researchers, scientists, and drug development professionals, it explores the foundational principles of LSERs, details their methodological application in predicting chemical leaching, addresses critical troubleshooting and optimization strategies to enhance model reliability, and presents rigorous validation and benchmarking against experimental data and alternative models. The synthesis of these aspects offers a validated, practical framework for employing LSERs in chemical safety risk assessments of pharmaceutical container-closure systems and medical devices, ultimately aiming to improve the accuracy of patient exposure estimations.
Understanding LSER Fundamentals and Their Role in Polymer-Water Partitioning
Core Principles of Linear Solvation Energy Relationships
Linear Solvation Energy Relationships (LSERs) are powerful predictive models used extensively in chemical, environmental, and pharmaceutical research. The fundamental premise of LSERs is that free-energy-related properties of a solute, such as partition coefficients, can be correlated through a linear relationship with molecular descriptors that characterize its interaction capabilities [1]. This approach allows researchers to predict a compound's behavior in different phases based on its molecular structure.
The Abraham solvation parameter model, a widely implemented form of LSER, utilizes two primary equations to quantify solute transfer between phases. For partitioning between two condensed phases, the model uses the equation:
log (P) = cp + epE + spS + apA + bpB + vpVx [1]
For gas-to-organic solvent partitioning, the relationship is expressed as:
log (KS) = ck + ekE + skS + akA + bkB + lkL [1]
In these equations, the lower-case coefficients (ep, sp, ap, bp, vp) represent system-specific parameters that describe the complementary effect of the solvent or phase on solute-solvent interactions. These coefficients are typically determined through multiple linear regression of experimental data and contain specific physicochemical information about the solvent system [1].
The Six Fundamental Solute Descriptors
The predictive power of LSER models stems from their use of six carefully defined solute descriptors that capture the key aspects of molecular interactions. The table below details these descriptors and their physicochemical significance.
Table 1: The Six Fundamental LSER Solute Descriptors
| Descriptor | Symbol | Molecular Interaction Represented | Determination Methods |
|---|---|---|---|
| Excess Molar Refraction | E | Polarizability from n- and π-electrons | Experimental via refractive index or Density Functional Theory (DFT) calculations [1] [2] [3] |
| Dipolarity/Polarizability | S | Dipolarity and overall polarizability | Calibrated QSPR models using theoretical molecular descriptors [2] [3] |
| Hydrogen Bond Acidity | A | Hydrogen bond donor ability (acidity) | Calibrated QSPR models using theoretical molecular descriptors [2] [3] |
| Hydrogen Bond Basicity | B | Hydrogen bond acceptor ability (basicity) | Calibrated QSPR models using theoretical molecular descriptors [2] [3] |
| McGowan's Characteristic Volume | Vx | Dispersion interactions and molecular size | Computed from molecular structure using DFT [1] [2] |
| Gas-Hexadecane Partition Coefficient | L | General dispersion interactions and hydrophobicity | Experimental or computed from DFT [1] [2] |
The E descriptor represents the solute's polarizability originating from its n- and π-electrons, while the S descriptor accounts for its dipolarity and polarizability. The A and B descriptors quantitatively express the solute's hydrogen-bonding potential as a donor and acceptor, respectively. The Vx descriptor characterizes the molecule's size and its capacity for dispersion interactions, and the L descriptor provides a measure of its hydrophobicity and general dispersive forces [1].
LSER Model for LDPE/Water Partitioning: Experimental Data and Performance
The application of LSERs to predict partitioning between low-density polyethylene (LDPE) and water is particularly relevant for assessing the leaching of compounds from plastic packaging into pharmaceutical products or food. A robust LSER model was recently developed and calibrated using experimental partition coefficients for 159 chemically diverse compounds [4] [5].
The established LSER model for the LDPE/water system is:
logKi,LDPE/W = -0.529 + 1.098E - 1.557S - 2.991A - 4.617B + 3.886V [6] [4] [5]
The performance of this model and a simpler log-linear model is summarized in the table below.
Table 2: Performance of LDPE/Water Partition Coefficient Prediction Models
| Model Type | Equation | Applicability | Statistics | Reference |
|---|---|---|---|---|
| LSER Model | logKi,LDPE/W = -0.529 + 1.098E - 1.557S - 2.991A - 4.617B + 3.886V | All compounds (polar & nonpolar) | n = 156, R² = 0.991, RMSE = 0.264 | [4] [5] |
| Log-Linear Model (Nonpolar Compounds Only) | logKi,LDPE/W = 1.18 logKi,O/W - 1.33 | Nonpolar compounds with low H-bonding propensity | n = 115, R² = 0.985, RMSE = 0.313 | [4] [5] |
| Log-Linear Model (All Compounds) | logKi,LDPE/W = 1.18 logKi,O/W - 1.33 | All compounds (including polar) | n = 156, R² = 0.930, RMSE = 0.742 | [4] [5] |
The model was further validated using an independent set of 52 compounds (约 33% of the total data). When using experimental solute descriptors for this validation set, the model maintained high performance (R² = 0.985, RMSE = 0.352). Even when using predicted solute descriptors from a QSPR tool, the model performed well (R² = 0.984, RMSE = 0.511), demonstrating its robustness for predicting partition coefficients for compounds with no experimental descriptor data [6].
Computational Protocols for Solute Descriptor Prediction
For high-throughput screening or when dealing with compounds for which experimental descriptor data is unavailable, in silico methods provide a valuable alternative. A comprehensive package solution has been developed to derive all necessary LSER solute parameters computationally [2] [3].
The workflow for this in silico derivation is as follows:
- Excess Molar Refraction (E), Molar Volume, and log L: These parameters are computed directly using Density Functional Theory (DFT) calculations [2] [3].
- Dipolarity/Polarizability (S), H-Bond Acidity (A), and Basicity (B): These are estimated by calibrated Quantitative Structure-Activity Relationship (QSPR) models that use theoretical molecular descriptors as inputs [2] [3].
LSER models constructed using these in silico solute parameter values have demonstrated performance comparable to conventional models that rely on empirical solute parameters, thereby greatly expanding the application domain of LSERs [2] [3].
Experimental Methodologies for Partition Coefficient Determination
Conventional Two-Phase Method
The traditional approach for determining polymer-water partition coefficients (KPE-w) involves allowing chemicals to reach equilibrium concentrations in direct contact polymer and water phases, followed by analytical measurement of both phases. However, this method presents significant challenges for highly hydrophobic organic compounds (HOCs), including low aqueous phase concentrations, long equilibration times (up to 365 days for some compounds), and potential analyte losses to surfaces [7].
Novel Three-Phase System
To overcome these limitations, a novel three-phase system has been developed that introduces a surfactant micellar pseudo-phase into the polymer/water system [7]. This method determines KPE-w indirectly from two more easily measurable parameters:
- Kmic-w: The micelle-water partition coefficient
- KPE-mic: The LDPE-micelle partition coefficient
The KPE-w value is then calculated from these two parameters. This approach sharply decreases the necessary equilibration time (to approximately half a month), avoids analytical challenges associated with direct water concentration measurements, provides concentrations far above analytical detection limits, and ultimately yields KPE-w values with smaller experimental errors [7].
Comparative Analysis with Other Polymer Phases
LSER system parameters enable direct comparison of sorption behavior across different polymer phases. When compared to polydimethylsiloxane (PDMS), polyacrylate (PA), and polyoxymethylene (POM), LDPE demonstrates distinct characteristics [6].
The heteroatomic building blocks in polymers like POM and PA provide capabilities for polar interactions, causing them to exhibit stronger sorption than LDPE for more polar, non-hydrophobic sorbates up to a logKi,LDPE/W range of 3 to 4. Above this range, all four polymers exhibit roughly similar sorption behavior for highly hydrophobic compounds [6].
Furthermore, by converting partition coefficients to logKi,LDPEamorph/W (considering only the amorphous fraction of the polymer as the effective phase volume), the resulting LSER model constant changes from -0.529 to -0.079, rendering the model more similar to a corresponding LSER model for n-hexadecane/water partitioning [6].
Essential Research Toolkit for LSER Applications
Table 3: Essential Research Reagents and Materials for LSER Studies
| Item | Specification/Example | Function in Research |
|---|---|---|
| Polymer Phase | Low-Density Polyethylene (LDPE), purified by solvent extraction | Primary sorbent phase for partitioning experiments; purification increases sorption capacity for polar compounds by up to 0.3 log units [4] [5]. |
| Surfactant | Polyoxyethylene (4) lauryl ether (Brij 30) | Forms micellar pseudo-phase in three-phase systems, enabling accurate determination of partition coefficients for HOCs [7]. |
| Reference Compounds | n-Hexadecane | Standard solvent for determining the L solute descriptor (gas-hexadecane partition coefficient) [1]. |
| Chemical Standards | 159+ chemically diverse compounds spanning wide MW (32-722) and polarity ranges (logK_{i,O/W}: -0.72 to 8.61) | Model calibration and validation; ensures coverage of relevant chemical space for leachables [4] [5]. |
| Computational Software | Density Functional Theory (DFT) packages, QSPR prediction tools | In silico derivation of solute descriptors (E, Vx, L, S, A, B) when experimental data is unavailable [2] [3]. |
The Critical Need for Predicting LDPE/Water Partitioning in Pharmaceutical Applications
In the pharmaceutical industry, the interaction between plastic materials and drug products is a critical safety consideration. When pharmaceutical solutions, such as intravenous drugs or liquid formulations, come into contact with low-density polyethylene (LDPE) packaging or delivery systems, organic compounds may leach from the polymer into the product, or drug components may partition into the polymer material. This exchange is principally governed by the equilibrium partition coefficient between the polymer and the aqueous phase (log Ki,LDPE/W), which dictates the maximum accumulation of a leachable and thus, patient exposure by leachables [5]. Accurate prediction of these partition coefficients is therefore fundamental to chemical safety risk assessments and ensuring patient safety, particularly for systems attaining equilibrium before the end of shelf-life.
The challenge for researchers and drug development professionals lies in obtaining reliable partition coefficients. Traditional experimental methods are often arduous, time-consuming, and prone to error, especially for highly hydrophobic compounds with low aqueous solubility which can require equilibration times as long as 365 days [7] [8]. This creates a pressing need for robust, accurate, and efficient predictive tools that can keep pace with modern drug development timelines while ensuring rigorous safety standards.
Experimental Approaches for Determining LDPE-Water Partition Coefficients
Conventional Methodological Challenges
Traditional methods for determining LDPE-water partition coefficients (KPE-w) typically involve allowing chemicals to reach equilibrium concentrations in polymer and water phases in direct contact with each other, followed by analysis of both phases [7]. This direct approach presents significant challenges for pharmaceutical applications, particularly with highly hydrophobic compounds which have correspondingly low aqueous phase concentrations and require exceptionally long equilibration times [7]. Furthermore, the analytical procedure itself is problematic because the water concentrations of contaminants of interest are often so small that processes like losses to surfaces can significantly impact results [7]. These limitations have driven the development of improved experimental and computational approaches.
Advancements in Experimental Methodology: The Three-Phase System
A novel three-phase partitioning system has been developed to address the limitations of conventional methods. This approach introduces a surfactant (Brij 30) to form a micellar pseudo-phase within the traditional polymer/water system [7] [9]. The method determines KPE-w values through a combination of two more easily and accurately measured values: the micelle-water partition coefficient (Kmic-w) and the LDPE-micelle partition coefficient (KPE-mic) [7].
The significant advantages of this system include a dramatic reduction in necessary equilibration time to approximately half a month, compared to much longer periods in traditional methods [7]. Additionally, it avoids the analytical challenges associated with directly measuring low aqueous phase concentrations, as the surfactant micelles enhance the solubility of hydrophobic compounds, bringing concentrations well above analytical detection limits [7] [9]. This method has been successfully applied to determine KPE-w values for a range of hydrophobic organic compounds, including polycyclic aromatic hydrocarbons (PAHs), polychlorinated biphenyls (PCBs), and polybrominated diphenyl ethers (PBDEs) with minimal experimental error [7].
Table 1: Key Experimental Methods for Determining LDPE-Water Partition Coefficients
| Method | Key Features | Advantages | Limitations | Suitable Compound Types |
|---|---|---|---|---|
| Traditional Two-Phase | Direct equilibrium measurement between LDPE and water phases [7] | Conceptually simple | Long equilibration times (up to 365 days); analytical challenges with low aqueous concentrations [7] | Compounds with moderate hydrophobicity |
| Three-Phase System (Micelle-Assisted) | Introduces surfactant micelles as a third pseudo-phase [7] [9] | Shorter equilibration (~15 days); avoids analytical challenges of low concentrations [7] | Requires additional calibration; surfactant purity critical | Hydrophobic organic compounds (PAHs, PCBs, PBDEs) [7] |
| Cosolvent Method | Uses polar organic solvents to enhance solubility [7] | Lowers polymer-liquid mixture partition coefficient | Nonlinear relationship may exist between chemical activities and cosolvent concentrations [7] | Limited range of polymer types |
The following workflow illustrates the experimental procedure for the novel three-phase partitioning system:
Computational Prediction Models for LDPE-Water Partitioning
Linear Solvation Energy Relationship (LSER) Models
Linear Solvation Energy Relationships (LSERs) represent a highly accurate and precise approach for estimating equilibrium partition coefficients involving polymeric phases. A robust LSER model for LDPE-water partitioning was recently calibrated using experimental data for 159 compounds spanning a wide range of chemical diversity, molecular weight, and polarity [5]. The model is expressed by the equation:
log Ki,LDPE/W = -0.529 + 1.098E - 1.557S - 2.991A - 4.617B + 3.886V [6] [10] [5]
This model has demonstrated exceptional performance with a coefficient of determination (R²) of 0.991 and root mean square error (RMSE) of 0.264 based on 156 observations [6] [5]. The capital letters in the equation represent Abraham solute descriptors: E (excess molar refraction), S (dipolarity/polarizability), A (hydrogen-bond acidity), B (hydrogen-bond basicity), and V (McGowan's molar volume) [6] [5]. The lowercase coefficients indicate the complementary properties of the partition system.
For pharmaceutical applications requiring estimation of partition coefficients for compounds without experimental LSER solute descriptors, predicted descriptors can be used with only a modest reduction in predictive performance (R² = 0.984, RMSE = 0.511) [6] [10]. This makes LSERs particularly valuable for screening potential leachables in early drug development stages.
Polyparameter Linear Free Energy Relationship (pp-LFER) and QSPR Models
Beyond LSERs, other modeling approaches have been developed for predicting LDPE-water partition coefficients. Polyparameter linear free energy relationship (pp-LFER) models have been established using descriptors such as McGowan's molar volume (V), hydrogen bond acceptor capacity (B), and hydrogen bond donor capacity (A) [8]. While these models provide valuable mechanistic insights, they sometimes exhibit lower correlation coefficients compared to comprehensive LSER models [8].
Quantitative Structure-Property Relationship (QSPR) models offer an alternative approach based solely on structural descriptors of chemicals. One such model developed using four influential descriptors (CrippenLogP, CIC0, MATS3i, and hydrogen bond donor capacity A) demonstrated good predictive capacity and robustness with R² values ranging from 0.771 to 0.921 [8] [11]. Mechanism interpretation from these models indicates that the factors influencing the distribution procedure between LDPE and water are primarily McGowan's molar volume and hydrophobic interactions [8].
Table 2: Comparison of Computational Models for Predicting LDPE-Water Partition Coefficients
| Model Type | Key Equation/Descriptors | Performance Metrics | Chemical Space | Pharmaceutical Application Context |
|---|---|---|---|---|
| LSER | log Ki,LDPE/W = -0.529 + 1.098E - 1.557S - 2.991A - 4.617B + 3.886V [6] [5] | R² = 0.991, RMSE = 0.264 (n=156) [5] | 159 compounds, wide chemical diversity [5] | Highest accuracy for safety assessments; suitable for diverse leachables |
| Log-Linear (nonpolar compounds) | log Ki,LDPE/W = 1.18 log Ki,O/W - 1.33 [5] | R² = 0.985, RMSE = 0.313 (n=115) [5] | Nonpolar compounds with low H-bonding propensity [5] | Useful for preliminary screening of nonpolar compounds |
| QSPR | CrippenLogP, CIC0, MATS3i, A (H-bond donor) [8] | R² = 0.771-0.921, Q² = 0.739-0.912 [8] | 268 compounds from 11 classes [8] | Broad applicability when experimental descriptors unavailable |
The conceptual framework below illustrates how these computational models interrelate and support predictive assessment of LDPE-water partitioning:
Essential Research Reagent Solutions for LDPE-Water Partitioning Studies
Successful experimental determination and modeling of LDPE-water partition coefficients requires specific, high-quality materials and computational resources. The following table details key research reagents and tools used in this field:
Table 3: Essential Research Reagents and Tools for LDPE-Water Partitioning Studies
| Reagent/Tool | Specifications | Function in Research | Application Context |
|---|---|---|---|
| Purified LDPE | Solvent-extracted to remove impurities [5] | Primary polymer phase for partitioning studies; purification critical for accurate measurements | Pharmaceutical packaging simulations; reference phase for passive sampling |
| Brij 30 Surfactant | Polyoxyethylene (4) lauryl ether; non-ionic [7] [9] | Forms micellar pseudo-phase in three-phase systems; enhances solubility of hydrophobic compounds | Enables measurement of highly hydrophobic compounds; reduces equilibration time |
| Abraham Solute Descriptors | E, S, A, B, V parameters [6] [5] | Input parameters for LSER models; describe molecular interactions | Predictive modeling of partition coefficients; chemical space characterization |
| Chemical Standards | PAHs, PCBs, PBDEs with purity >99% [7] | Model compounds for method validation and calibration | Establishing reference partition coefficients; model training and validation |
| QSPR Prediction Tools | Software for predicting LSER descriptors [6] [10] | Generates solute descriptors from chemical structure | Extending predictions to compounds without experimental descriptors |
Comparative Analysis of Polymer Sorption Behaviors
Understanding how LDPE compares to other common polymeric materials in pharmaceutical applications is essential for appropriate material selection. LSER system parameters enable direct comparison of sorption behaviors across different polymers:
LDPE exhibits distinctly different sorption characteristics compared to polymers like polydimethylsiloxane (PDMS), polyacrylate (PA), and polyoxymethylene (POM) [6]. The latter polymers, by offering capabilities for polar interactions due to their heteroatomic building blocks, exhibit stronger sorption than LDPE for more polar, non-hydrophobic compounds in the log Ki,LDPE/W range of 3 to 4 [6]. Above this range, all four polymers demonstrate roughly similar sorption behavior [6].
Notably, the sorption of polar compounds into pristine (non-purified) LDPE was found to be up to 0.3 log units lower than into purified LDPE [5]. This finding has significant implications for pharmaceutical testing protocols, as the purification state of the polymer can substantially impact partitioning behavior and thus, worst-case leaching assessments.
The accurate prediction of LDPE-water partition coefficients represents a critical capability in pharmaceutical development, directly impacting the assessment of leachable compounds and patient safety. While traditional experimental methods face challenges of prolonged equilibration times and analytical limitations, recent advancements in three-phase experimental systems and computational modeling have significantly enhanced our capabilities in this field.
The development of highly accurate LSER models, along with complementary pp-LFER and QSPR approaches, provides researchers with a robust toolkit for predicting partition coefficients across a diverse chemical space. These tools enable more reliable risk assessments of packaging systems and delivery devices, supporting the pharmaceutical industry's mission to ensure product safety and efficacy. As these predictive methodologies continue to evolve and validate against experimental data, they offer the promise of more efficient drug development processes without compromising scientific rigor or patient safety.
Linear Solvation Energy Relationships (LSERs) represent a powerful and widely adopted quantitative approach for predicting the partitioning behavior of solutes between different phases. Developed primarily by Abraham and coworkers, the LSER model provides a mechanistic framework for understanding and quantifying the intermolecular interactions that govern solvation and partitioning processes [12]. The model's robustness stems from its ability to deconstruct complex physicochemical phenomena into contributions from distinct, interpretable molecular interactions. This article provides a comprehensive term-by-term deconstruction of the fundamental LSER equation, with a specific focus on validating its predictions for Low-Density Polyethylene (LDPE)/water partitioning—a system of critical importance in pharmaceutical development and environmental chemistry.
The core LSER model for solute partitioning between two condensed phases is expressed as:
[ \log(P) = cp + epE + spS + apA + bpB + vpV_x ]
In this equation, (P) represents a partition coefficient, while the capital letters ((E), (S), (A), (B), (Vx)) denote solute-specific molecular descriptors. Conversely, the lower-case letters ((cp), (ep), (sp), (ap), (bp), (v_p)) are system-specific coefficients that characterize the complementary properties of the phases between which partitioning occurs [1] [12]. The model's predictive power lies in its linear free-energy relationship foundation, which allows for the extrapolation of a solute's behavior across different chemical systems once its descriptors are known.
Term-by-Term Analysis of the LSER Equation
The Constant Term ((c_p))
The constant term (c_p) serves as the regression intercept. Its value is system-dependent and can be influenced by factors such as the reference states used for the partition coefficient and properties inherent to the partitioning system that are not fully captured by the other five interaction terms [6]. For instance, in the LDPE/water system, the constant was determined to be -0.529 [6] [5]. When the partitioning is conceptually adjusted to consider only the amorphous fraction of LDPE as the effective volume, this constant shifts to -0.079, making the system's behavior more analogous to partitioning into a liquid phase like n-hexadecane [6]. This adjustment highlights how the constant term can absorb nuances related to phase morphology.
Excess Molar Refraction ((e_pE))
- Solute Descriptor ((E)): This parameter quantifies the solute's polarizability arising from π- and n-electrons. It is typically determined based on the solute's refractive index [12].
- System Coefficient ((e_p)): This coefficient reflects the capability of the system's phases to interact with a solute via polarizability-based interactions, such as π-π and n-π interactions [12].
- LDPE/Water Context: In the specific LSER model for LDPE/water partitioning, the system coefficient for the (E) term is positive (+1.098) [6] [5]. This indicates that solutes with higher polarizability (larger (E)) have a greater tendency to partition into the LDPE phase relative to water, as the dispersion forces favored by the polymer are more significant for polarizable molecules.
Dipolarity/Polarizability ((s_pS))
- Solute Descriptor ((S)): This descriptor represents the solute's dipolarity and polarizability. It is a measure of a molecule's ability to engage in dipole-dipole and dipole-induced dipole interactions [12].
- System Coefficient ((s_p)): This coefficient indicates the system's overall responsiveness to a solute's dipolarity and polarizability.
- LDPE/Water Context: The coefficient for this term in the LDPE/water system is negative (-1.557) [6] [5]. This strongly suggests that the aqueous phase competes more effectively than the LDPE phase for interactions with dipolar molecules. A solute with high dipolarity is therefore "penalized" in its partitioning to the non-polar polymer environment.
Hydrogen Bond Acidity ((a_pA))
- Solute Descriptor ((A)): This parameter quantifies the solute's hydrogen-bond donating ability (acidity) [12].
- System Coefficient ((a_p)): This coefficient reflects the difference in hydrogen-bond basicity between the two partitioning phases. A large negative value indicates that one phase is a much stronger hydrogen-bond acceptor than the other.
- LDPE/Water Context: The highly negative coefficient for this term (-2.991) underscores water's superior strength as a hydrogen-bond acceptor compared to LDPE [6] [5]. Solutes that are strong hydrogen-bond donors (high (A)) are strongly retained in the aqueous phase, making this one of the most significant terms for discouraging partitioning into LDPE.
Hydrogen Bond Basicity ((b_pB))
- Solute Descriptor ((B)): This descriptor represents the solute's hydrogen-bond accepting ability (basicity) [12].
- System Coefficient ((b_p)): This coefficient reflects the difference in hydrogen-bond acidity between the two phases.
- LDPE/Water Context: This term has the largest magnitude coefficient in the LDPE/water equation (-4.617) [6] [5]. This emphatically confirms that water's powerful ability to donate hydrogen bonds is a dominant interaction. Solutes with any significant hydrogen-bond accepting character (high (B)) will exhibit a very strong preference for the aqueous environment over the inert LDPE.
McGowan's Characteristic Volume ((vpVx))
- Solute Descriptor ((V_x)): This is McGowan's characteristic volume, which is calculated from molecular structure and represents the molecular size of the solute [1] [12].
- System Coefficient ((v_p)): This term is generally related to the cavity formation energy required to accommodate the solute within a solvent phase. It often correlates with the difference in cohesion of the two phases.
- LDPE/Water Context: The coefficient for this term is positive and has the second-largest magnitude (+3.886) [6] [5]. Creating a cavity in highly cohesive water is energetically far more costly than in the hydrophobic, low-cohesion LDPE. Consequently, larger solutes experience a significant thermodynamic driving force to partition into the polymer phase to minimize the disruptive cavity effect in water.
Table 1: Summary of Solute Descriptors in the LSER Model
| Term | Symbol | Molecular Interaction Property | Determination Basis |
|---|---|---|---|
| Excess Molar Refraction | (E) | Polarizability from π- and n-electrons | Refractive Index [12] |
| Dipolarity/Polarizability | (S) | Dipole-dipole and dipole-induced dipole interactions | Experimental polarity scales [12] |
| Hydrogen Bond Acidity | (A) | Hydrogen-bond donating ability | Experimental HB acidity scales [12] |
| Hydrogen Bond Basicity | (B) | Hydrogen-bond accepting ability | Experimental HB basicity scales [12] |
| McGowan's Volume | (V_x) | Molecular size | Molecular structure [1] |
Table 2: System Coefficients for LDPE/Water Partitioning and Their Chemical Interpretation [6] [5]
| System Coefficient | Value | Chemical Interpretation for LDPE/Water |
|---|---|---|
| (c_p) | -0.529 | Regression intercept; adjusts for system-specific references. |
| (e_p) | +1.098 | LDPE favors polarizable solutes more than water does. |
| (s_p) | -1.557 | Water competes more effectively for dipolar interactions. |
| (a_p) | -2.991 | Water is a much stronger hydrogen-bond acceptor than LDPE. |
| (b_p) | -4.617 | Water is a much stronger hydrogen-bond donor than LDPE. |
| (v_p) | +3.886 | Cavity formation is far more costly in water than in LDPE. |
LSER Model Validation in LDPE/Water Partitioning
Experimental Protocols for Model Calibration
The validation and calibration of an LSER model for a specific system like LDPE/water require a rigorous experimental protocol. A representative methodology, as employed in recent robust studies, involves the following key steps [6] [5]:
- Material Preparation: Low-density polyethylene film is often purified via solvent extraction (e.g., using ethanol and n-pentane) to remove manufacturing additives and impurities that could interfere with sorption measurements. Both purified and pristine (non-purified) LDPE can be studied to quantify the effect of purification.
- Solute Selection: A large set of chemically diverse compounds (e.g., 150+ solutes) is selected to ensure the model is trained across a wide chemical space. The compounds should span a broad range of molecular weight, hydrophobicity (log (K_{i,O/W})), vapor pressure, and polarity, with special attention to varying hydrogen-bond donor and acceptor capacities.
- Partition Coefficient Determination: Experimental partition coefficients ((K_{i,LDPE/W})) are determined by measuring the equilibrium distribution of each solute between the LDPE film and an aqueous buffer solution. Analysis of solute concentrations in both phases is typically performed using chromatographic methods (e.g., HPLC or GC-MS).
- Data Compilation and Regression: Experimentally determined log (K{i,LDPE/W}) values are compiled for all solutes. Using multiple linear regression analysis, the log (K{i,LDPE/W}) values are fitted against the known solute descriptors ((E, S, A, B, Vx)) for the training set, yielding the system-specific coefficients ((cp, ep, sp, ap, bp, v_p)).
Model Performance and Benchmarking
The calibrated LSER model for LDPE/water partitioning has demonstrated exceptional predictive performance. The model for purified LDPE, based on 156 compounds, was reported with a coefficient of determination (R²) of 0.991 and a root mean square error (RMSE) of 0.264 log units [6] [5]. When validated on an independent set of 52 compounds using experimental solute descriptors, the model maintained high performance (R² = 0.985, RMSE = 0.352) [6]. Even when solute descriptors were predicted in silico rather than measured experimentally, the model remained robust (R² = 0.984, RMSE = 0.511), which is crucial for predicting the behavior of compounds for which experimental descriptors are unavailable [6].
Comparison with Other Polymers and Phases
LSER system parameters enable direct comparison of the sorption behavior of LDPE with other polymers. For instance, when compared to polydimethylsiloxane (PDMS), polyacrylate (PA), and polyoxymethylene (POM), LDPE exhibits the weakest capacity for polar interactions due to its lack of heteroatomic building blocks [6]. Consequently, for polar, non-hydrophobic solutes (with log (K{i,LDPE/W}) up to 3-4), PA and POM show stronger sorption than LDPE. However, for very hydrophobic compounds (log (K{i,LDPE/W}) > 4), all four polymers exhibit roughly similar sorption behavior, as hydrophobic effects dominate [6].
Diagram 1: LSER Model Validation Workflow. This flowchart outlines the key experimental and computational steps for calibrating and validating an LSER model for LDPE/water partitioning.
The Scientist's Toolkit: Essential Research Reagents and Materials
Table 3: Essential Research Reagents and Materials for LSER and Partitioning Studies
| Item | Function / Relevance in LSER Research |
|---|---|
| Purified LDPE Film | The polymer phase of interest; purification via solvent extraction is critical to remove interfering additives for accurate partition coefficient measurement [5]. |
| Chemical Standards | A diverse set of solute compounds with pre-established Abraham LSER descriptors ((E, S, A, B, V_x)), essential for model calibration and validation [6] [5]. |
| Aqueous Buffer Solutions | The aqueous partitioning phase; buffers ensure consistent pH, which can influence the ionization state and partitioning of ionizable solutes. |
| LSER Database | A freely accessible, curated database containing solute descriptors and system coefficients, enabling the outright calculation of partition coefficients [6]. |
| Chromatography Systems (HPLC, GC-MS) | Essential analytical tools for quantifying solute concentrations in both the aqueous and polymer phases after equilibrium is reached [5]. |
| QSPR Prediction Software | In-silico tools for predicting Abraham solute descriptors for novel compounds when experimental determination is not feasible [6]. |
The term-by-term deconstruction of the LSER equation reveals a model of remarkable physicochemical intuition and predictive power. The analysis of the LDPE/water system coefficients provides a clear, quantitative narrative: partitioning from water into the polymer is overwhelmingly driven by the solute's size ((V_x)) and polarizability ((E)), which favor the LDPE phase, and is strongly opposed by the solute's hydrogen-bonding capacity ((A, B)) and dipolarity ((S)), which favor the aqueous phase. The exceptionally large negative coefficients for the hydrogen-bonding terms quantitatively confirm the dominance of these interactions in retaining polar solutes in water. The robust experimental validation of the LSER model for LDPE/water partitioning, achieving high precision across a vast chemical space, solidifies its status as an indispensable tool for accurate risk assessment in pharmaceutical development and beyond.
In pharmaceutical development, extractables and leachables (E&L) represent a critical quality attribute for container-closure systems, manufacturing components, and drug delivery devices. Extractables are chemical compounds that can be released from materials under exaggerated conditions (e.g., extreme temperatures, aggressive solvents), while leachables are those that migrate under normal product storage and use conditions, potentially ending up in the final drug product [13]. The chemical space encompassing potential leachables is vast, covering compounds with diverse molecular weights, polarities, and functional groups, from nonpolar polymer additives to highly polar degradation products [5]. Understanding and predicting the behavior of compounds across this chemical space is fundamental to assessing patient exposure and conducting accurate toxicological risk assessments.
A pivotal parameter in this assessment is the partition coefficient, which quantifies the distribution of a compound between two phases at equilibrium. For pharmaceutical systems utilizing low-density polyethylene (LDPE) and other polymers, the LDPE-water partition coefficient (K_{i,LDPE/W}) directly influences the maximum possible accumulation of a leachable and, consequently, patient exposure [5]. Traditional predictive modeling often relied on coarse estimations or simplistic log-linear relationships with octanol-water partitioning, but these approaches frequently failed for polar compounds, creating significant gaps in chemical safety risk assessments [5] [14]. This guide evaluates the performance of Linear Solvation Energy Relationships (LSERs) against traditional models in predicting LDPE-water partitioning, providing researchers with validated experimental protocols and data-driven comparisons.
Model Comparison: LSERs vs. Traditional Approaches
Theoretical Foundations
Linear Solvation Energy Relationships (LSERs) are multi-parameter models that describe partitioning behavior based on a compound's specific molecular interactions. The general LSER form for the LDPE-water system is expressed as [5]: [ \log K_{i,LDPE/W} = c + eE + sS + aA + bB + vV ] where the capital letters represent solute descriptors (E for excess molar refractivity, S for dipolarity/polarizability, A for hydrogen-bond acidity, B for hydrogen-bond basicity, and V for McGowan's characteristic volume), and the lowercase letters are system-specific coefficients that are calibrated against experimental data.
In contrast, the traditional log-linear model relies on a simplistic correlation with the octanol-water partition coefficient (K{i,O/W}) [5]: [ \log K{i,LDPE/W} = m \log K_{i,O/W} + c ] This model assumes that partitioning is dominated by hydrophobicity, an assumption that holds reasonably well for nonpolar compounds but breaks down significantly for mono- and bipolar molecules.
Performance Evaluation and Comparative Data
Recent research has provided robust, quantitative comparisons of these models' performances. The table below summarizes key findings from a comprehensive study that calibrated both models using a chemically diverse set of 156 compounds [5].
Table 1: Model Performance Comparison for Predicting LDPE-Water Partitioning
| Model Type | Chemical Scope | Sample Size (n) | Coefficient of Determination (R²) | Root Mean Square Error (RMSE) | Key Limitations |
|---|---|---|---|---|---|
| LSER Model | Wide range, including polar compounds | 156 | 0.991 | 0.264 | Requires a full set of five Abraham solute descriptors |
| Log-Linear Model (Nonpolar Compounds Only) | Nonpolar, low H-bonding propensity | 115 | 0.985 | 0.313 | Fails for polar chemicals; limited applicability |
| Log-Linear Model (Full Compound Set) | Includes polar compounds | 156 | 0.930 | 0.742 | Poor predictive power for hydrogen-bonding and polar molecules |
The data demonstrates the clear superiority of the LSER model across a broad chemical space. Its high R² and low RMSE indicate it is both accurate and precise. The log-linear model shows utility for a specific, narrow band of the chemical space (nonpolar compounds) but becomes unreliable when applied to the wider universe of potential leachables, as evidenced by the R² dropping to 0.93 and the RMSE increasing nearly three-fold [5]. This performance gap is critical because polar leachables, which may arise from interactions between a drug formulation and a polymer, are not uncommon and can pose significant toxicological risks [13].
Experimental Protocols for Model Validation
Validating predictive models requires robust experimental data. The following sections detail two advanced methodologies for determining LDPE-water partition coefficients, which can be used to calibrate and verify LSER and other model predictions.
Direct Measurement via a Novel Three-Phase System
Objective: To accurately determine LDPE-water partition coefficients (K_PE-w) for hydrophobic organic compounds, overcoming challenges like long equilibration times and low aqueous concentrations [7].
Workflow:
- Equilibration: The target compound is allowed to equilibrate between LDPE, an aqueous phase, and a micellar pseudo-phase created by adding a surfactant like Brij 30.
- Measurement of K_PE-mic: The partition coefficient between LDPE and the micelle is measured. Concentrations are high in both organic phases, minimizing analytical error.
- Measurement of K_mic-w: The partition coefficient between the micelle and water is measured. The surfactant enhances solubility, making determination easier.
- Calculation of KPE-w: The LDPE-water partition coefficient is calculated from the two measured values: KPE-w = KPE-mic × Kmic-w.
This method reduces equilibration time to approximately half a month and avoids the analytical difficulties of measuring very low aqueous concentrations directly [7].
Cosolvency Approach for Simulating Clinical Conditions
Objective: To predict solute partitioning between LDPE and water-ethanol mixtures that simulate the polarity of clinically relevant media, enabling more realistic patient exposure estimates [14].
Workflow:
- Solubility Calculation: The hypothetical partition coefficient based on solubility in water-ethanol mixtures versus pure water is calculated using a cosolvency model. The LSER-based cosolvency model has been shown to be slightly superior to the log-linear model of Yalkowsky and coworkers [14].
- Thermodynamic Cycle Application: Using a thermodynamic cycle and a pre-existing LSER model for the LDPE-water partition coefficient, the partitioning between LDPE and the ethanol-water mixture is calculated.
- Experimental Verification: The calculated values are experimentally verified for a wide array of chemically diverse solutes in ethanol-water mixtures with volume fractions of 0.1, 0.2, 0.35, and 0.5. The LSER-based approach showed good correlation with experimentally obtained values [14].
This protocol allows for the tailored preparation of simulating solvents based on the clinical medium, improving the reliability of risk assessments.
The following diagram illustrates the logical workflow for the cosolvency model approach to predicting partitioning in simulating solvents.
Figure 1: Workflow for the cosolvency model approach to predicting partitioning.
The Scientist's Toolkit: Essential Research Reagents and Materials
Successful experimental execution in E&L partitioning research relies on specific, high-quality materials. The following table catalogues key reagents and their critical functions in the protocols described above.
Table 2: Essential Research Reagents for LDPE-Water Partitioning Studies
| Reagent/Material | Specifications | Function in Experiment | Key Considerations |
|---|---|---|---|
| Low-Density Polyethylene (LDPE) | Purified by solvent extraction; defined thickness | Primary polymer phase for partitioning | Pristine vs. purified LDPE can show differing sorption for polar compounds [5] |
| Chemical Standards | 159+ compounds spanning a wide chemical space (MW: 32-722, log K_i,O/W: -0.72 to 8.61) [5] | Model solutes for calibrating and validating models | Must include nonpolar, monopolar, and bipolar compounds to represent the "universe" of leachables [5] |
| Surfactant (Brij 30) | Polyoxyethylene (4) lauryl ether, non-ionic | Forms micellar pseudo-phase in three-phase system | Low hydrophile-lipophile balance number increases efficiency; reduces equilibration time [7] |
| Simulating Solvents (Ethanol-Water Mixtures) | Binary mixtures with defined volume fractions (e.g., 10-50% ethanol) [14] | Mimic the extraction strength of clinically relevant media | Allows for tailored polarity to match specific drug products for realistic exposure estimates [14] |
| Abraham Solute Descriptors | Dataset for E, S, A, B, V descriptors for all test compounds | Core input parameters for LSER model calibration | Accuracy and availability of descriptors can be a limiting factor for model application |
The move towards risk-based approaches in regulatory science for E&L testing demands more sophisticated and accurate predictive tools [15]. Relying on coarse estimations or models with limited applicability to polar chemicals is no longer sufficient. The evidence clearly indicates that LSER models provide a robust, accurate framework for predicting LDPE-water partitioning across a diverse chemical space, directly supporting more reliable safety risk assessments.
As regulatory landscapes evolve toward greater global harmonization, the emphasis on comprehensive toxicological data and robust analytical validation will only intensify [15]. Integrating high-performance predictive models like LSERs into the chemical assessment triad—which encompasses controlled extraction studies, leachable testing, and toxicological evaluation—represents a best practice for the pharmaceutical industry. This approach not only ensures patient safety but also enhances development efficiency by enabling more targeted and relevant experimental studies.
Equilibrium Partitioning as the Driver for Leachable Accumulation
In the pharmaceutical and food industries, accurately predicting the accumulation of leachable substances from plastic packaging into drug products is a critical aspect of chemical safety risk assessments. When equilibrium between a plastic material and a liquid product is reached within the product's shelf-life, the partition coefficient governing the distribution of a substance between the polymer and solution phases becomes the principal factor dictating maximum patient exposure to leachables. Despite its fundamental importance, the accurate prediction of these partition coefficients, particularly for low-density polyethylene (LDPE)—one of the most common packaging materials—has remained challenging. Traditional predictive approaches often rely on coarse estimations, lacking the precision required for robust safety assessments. This guide objectively compares the performance of contemporary experimental and in silico methodologies for determining LDPE-water partition coefficients (KPE-W), with particular emphasis on validating Linear Solvation Energy Relationship (LSER) predictions within a comprehensive research framework. By comparing the technical principles, experimental requirements, and predictive performance of each approach, we provide drug development professionals with evidence-based guidance for selecting appropriate methodologies based on their specific assessment needs.
Methodological Comparison: Experimental & Computational Approaches
Experimental Methodologies for Partition Coefficient Determination
Conventional Two-Phase Equilibrium Method
The conventional approach for determining polymer-water partition coefficients involves allowing chemicals to reach equilibrium concentrations between polymer and water phases, followed by analytical measurement of both phases [7]. While conceptually straightforward, this method presents significant practical challenges, particularly for highly hydrophobic organic compounds (HOCs) with limited water solubility [7]. These challenges include:
- Extended equilibration times: Studies report sampling periods as long as 365 days for polybrominated diphenyl ethers (PBDEs) [7]
- Analytical limitations: Direct measurement of trace-level aqueous phase concentrations approaches or falls below analytical detection limits for many HOCs [7]
- Experimental artifacts: Potential losses to surfaces and other systematic errors affect measurement accuracy [7]
The cosolvent method represents an alternative approach that mitigates some solubility limitations by using polar organic solvents (methanol, acetone) to enhance compound solubility [7]. Through linear extrapolation to 0% cosolvent, researchers can estimate polymer-water partition coefficients. However, this method can yield inaccurate results due to potential nonlinear relationships between chemical activities and cosolvent concentrations [7]. Additionally, it is labor-intensive and applicable only to a limited range of polymer types [7].
Novel Three-Phase Partitioning System
A recently developed three-phase system addresses several limitations of conventional methods by introducing a surfactant micellar pseudo-phase into the traditional polymer/water system [7]. This innovative approach involves:
- System composition: Addition of sufficient surfactant (Brij 30) to form micelles within the LDPE/water system [7]
- Indirect determination: KPE-W values are derived from two experimentally measured parameters—the micelle-water partition coefficient (Kmic-w) and the LDPE-micelle partition coefficient (KPE-mic) [7]
- Mathematical relationship: KPE-W = KPE-mic × Kmic-w [7]
This methodology significantly reduces equilibration time to approximately half a month while avoiding analytical challenges associated with direct aqueous phase concentration measurements [7]. The approach maintains measurement precision even for highly hydrophobic compounds because concentrations in both organic phases (polymer and micellar) remain well above analytical detection limits [7].
Rapid Vacuum-Based Method for Surface-Air Partitioning
While not directly applicable to LDPE-water systems, a novel vacuum-based chamber approach has been developed for rapidly determining surface-air partition coefficients of semivolatile organic compounds (SVOCs) [16]. This method accelerates sorption equilibration from weeks to hours by reducing boundary-layer resistance under low-pressure conditions [16]. The demonstrated principle that pressure variations do not affect partition coefficient values while dramatically reducing equilibration time may inspire future innovations in polymer-water partitioning methodologies.
Table 1: Comparison of Experimental Methods for Determining Partition Coefficients
| Method | Time Requirements | Applicable Compound Range | Key Advantages | Key Limitations |
|---|---|---|---|---|
| Conventional Two-Phase | Weeks to months (up to 365 days) | Limited for highly hydrophobic compounds | Conceptually straightforward; direct measurement | Long equilibration; analytical challenges at low concentrations |
| Cosolvent Method | Moderate to high | Limited by cosolvent interactions | Improves solubility for challenging compounds | Potential nonlinear behavior; labor-intensive; limited polymer applicability |
| Three-Phase System | Approximately 15 days | Broad range, including HOCs | Reduced equilibration time; avoids direct aqueous phase measurement | Requires additional partition measurements; surfactant introduction |
| Vacuum-Based (air) | Hours | SVOCs for surface-air partitioning | Extremely rapid equilibration; high-throughput capability | Currently limited to surface-air systems |
In Silico Predictive Models
Linear Solvation Energy Relationships (LSERs)
LSERs represent a highly accurate approach for predicting partition coefficients between LDPE and water. A robust LSER model was recently developed based on an extensive dataset of 159 compounds spanning wide molecular weight (32-722 g/mol), octanol-water partition coefficient (log Ki,O/W: -0.72 to 8.61), and LDPE-water partition coefficient (log Ki,LDPE/W: -3.35 to 8.36) ranges [5]. The calibrated model for purified LDPE material is expressed as:
log Ki,LDPE/W = -0.529 + 1.098E - 1.557S - 2.991A - 4.617B + 3.886V [5]
Where E represents the excess molar refractivity, S represents dipolarity/polarizability, A and B represent hydrogen-bond acidity and basicity, and V represents the McGowan characteristic volume [5]. This model demonstrates exceptional accuracy and precision (n = 156, R² = 0.991, RMSE = 0.264) across a chemically diverse compound space representative of potential leachables from pharmaceutical plastics [5].
Independent validation using approximately 33% of the total observations (n = 52) as an external validation set confirmed the model's predictive power, yielding R² = 0.985 and RMSE = 0.352 when using experimental solute descriptors [10]. When employing QSPR-predicted descriptors instead of experimental ones, the statistics remained strong (R² = 0.984, RMSE = 0.511), indicating utility for compounds lacking experimentally determined descriptors [10].
Log-Linear Octanol-Water Correlation
For nonpolar compounds with low hydrogen-bonding donor and/or acceptor propensity, a log-linear correlation against octanol-water partition coefficients provides reasonable estimates [5]. The model for nonpolar compounds is:
log Ki,LDPE/W = 1.18 log Ki,O/W - 1.33 (n = 115, R² = 0.985, RMSE = 0.313) [5]
However, this approach shows significantly reduced performance when applied to mono-/bipolar compounds (n = 156, R² = 0.930, RMSE = 0.742), rendering it of limited value for polar compounds commonly encountered in pharmaceutical applications [5].
Quantitative Structure-Property Relationships (QSPRs)
Several QSPR software packages are available for predicting partition coefficients and other physicochemical properties relevant to leachable accumulation assessments. A recent comparative analysis evaluated IFSQSAR, OPERA, and EPI Suite for prediction accuracy, applicability domain, and uncertainty [17]. Key findings include:
- Uncertainty characterization: IFSQSAR's 95% prediction interval (PI95) calculated from root mean squared error of prediction (RMSEP) captured 90% of external experimental data, while OPERA and EPI Suite required factor increases of at least 4 and 2 respectively for their PI95 to achieve similar coverage [17]
- Data-poor chemicals: Polyfluorinated alkyl substances (PFAS), ionizable organic chemicals (particularly strong acids and bases), and complex multifunctional structures represent significant challenges for all predictive approaches [17]
- Consensus predictions: Combining predictions from multiple models can provide more reliable estimates, though methodological differences must be considered [17]
Table 2: Comparison of In Silico Predictive Models for LDPE-Water Partition Coefficients
| Model Type | Theoretical Basis | Applicability Domain | Performance Metrics | Implementation Requirements |
|---|---|---|---|---|
| LSER | Linear free-energy relationships based on solute descriptors | Broad chemical diversity | R² = 0.991, RMSE = 0.264 (calibration) [5] | Experimental or predicted solute descriptors |
| Log-Linear Correlation | Empirical correlation with octanol-water partitioning | Limited to nonpolar compounds | R² = 0.985, RMSE = 0.313 (nonpolar compounds only) [5] | Octanol-water partition coefficient data |
| QSPR (IFSQSAR) | Structural descriptors and machine learning | Defined applicability domain with uncertainty quantification | Captures 90% of external data with native PI95 [17] | Chemical structure input |
| QSPR (OPERA) | Structural descriptors and machine learning | Defined applicability domain | Requires ~4x PI95 increase to capture 90% of external data [17] | Chemical structure input |
| QSPR (EPI Suite) | Structural fragments and group contribution methods | Limited explicit applicability domain | Requires ~2x PI95 increase to capture 90% of external data [17] | Chemical structure input |
Experimental Protocols & Methodologies
Three-Phase Partitioning System Protocol
The novel three-phase partitioning method provides a robust experimental approach for determining LDPE-water partition coefficients while overcoming traditional methodological limitations [7].
Materials and Reagents:
- LDPE films: Purified through solvent extraction to remove interfering additives [5]
- Surfactant: Polyoxyethylene (4) lauryl ether (Brij 30) with low hydrophile-lipophile balance number [7]
- Target compounds: Hydrophobic organic compounds (HOCs) including PAHs, PBDEs, and PCBs [7]
- Analytical standards: Deuterated or otherwise distinguishable reference compounds for quantification [7]
Experimental Procedure:
- Micelle-water partitioning: Determine critical micelle concentration (CMC) of Brij 30 using surface tension measurements [7]
- Solubility enhancement measurement: Measure total chemical solubility in Brij 30 solutions above CMC; plot concentration in micellar pseudo-phase (Cm) versus surfactant concentration (Xm) [7]
- Kmic-w calculation: Calculate micelle-water partition coefficient from slope of solubility enhancement curve [7]
- LDPE-micelle partitioning: Equilibrate LDPE strips with chemical solutions in Brij 30; measure concentrations in both phases after equilibration [7]
- KPE-mic calculation: Determine LDPE-micelle partition coefficient from concentration ratio in polymer and micellar phases [7]
- KPE-w determination: Calculate LDPE-water partition coefficient as product of KPE-mic and Kmic-w [7]
Validation: The method demonstrated strong correlation with conventional measurements for reference compounds (R² = 0.897, p < 0.001) while providing improved precision, particularly for highly hydrophobic compounds [7].
Direct LDPE-Water Partitioning Measurement
For compounds amenable to traditional measurement, the direct equilibrium method remains valuable, particularly when conducted with purified LDPE [5].
Materials Preparation:
- LDPE purification: Extract pristine LDPE films with organic solvents (e.g., methanol, hexane) to remove additives and impurities [5]
- Aqueous buffer preparation: Use clinically relevant media or standardized buffers; consider sterilization for pharmaceutical relevance [5]
- Chemical dosing: Prepare stock solutions in appropriate solvents; verify concentration and purity [5]
Equilibration and Sampling:
- System setup: Place purified LDPE films in contact with aqueous solutions containing target compounds; include appropriate controls [5]
- Equilibration: Maintain systems at constant temperature with agitation; monitor approach to equilibrium through time-series sampling [5]
- Phase separation: Separate polymer and aqueous phases after equilibration; minimize disturbance of established equilibrium [5]
- Extraction and analysis: Extract compounds from both phases; use appropriate internal standards; employ sensitive analytical methods (GC-MS, LC-MS) [5]
Data Interpretation:
- Partition coefficient calculation: Determine KPE-W as ratio of equilibrium concentrations in polymer and aqueous phases [5]
- Quality control: Assess mass balance to identify potential experimental artifacts; verify equilibrium attainment through time-course studies [5]
Visualization of Methodological Approaches
Experimental Workflow Comparison
Diagram 1: Method Selection Workflow for LDPE-Water Partition Coefficient Determination
LSER Model Development and Validation Pathway
Diagram 2: LSER Model Development and Validation Pathway
The Scientist's Toolkit: Essential Research Reagents & Materials
Table 3: Essential Research Reagents and Materials for LDPE-Water Partitioning Studies
| Item | Specification/Example | Function/Purpose | Key Considerations |
|---|---|---|---|
| LDPE Material | Purified grade, solvent-extracted | Primary polymer phase for partitioning studies | Purity affects sorption behavior; purified LDPE shows different partitioning for polar compounds [5] |
| Surfactant | Brij 30 (Polyoxyethylene (4) lauryl ether) | Forms micellar pseudo-phase in three-phase systems | Low HLB number enhances efficiency for HOCs; must determine CMC [7] |
| Chemical Standards | PAHs, PCBs, PBDEs, pharmaceutical relevant compounds | Model compounds for method development and validation | Purity >99%; include deuterated analogs as internal standards [7] |
| Solvent Systems | Methanol, ethanol, hexane for extraction and cleaning | LDPE purification and compound extraction | Purity affects background interference; residue analysis recommended [5] |
| Aqueous Media | Phosphate buffers, simulated physiological solutions | Clinically relevant contact media for partitioning | pH and ionic strength affect partitioning; sterilization may be required [5] |
| LSER Descriptors | Experimental or predicted E, S, A, B, V parameters | Input variables for LSER predictions | Experimental descriptors preferred; QSPR-predicted descriptors increase uncertainty [10] |
| QSPR Software | IFSQSAR, OPERA, EPI Suite | In silico prediction of partition coefficients | Differ in applicability domain and uncertainty characterization [17] |
Performance Benchmarking & Comparative Analysis
Accuracy and Precision Metrics
The evaluated methodologies demonstrate distinct performance characteristics in predicting and measuring LDPE-water partition coefficients:
Experimental Methods:
- Three-phase system: Shows strong correlation with conventional measurements (R² = 0.897, p < 0.001) while significantly reducing equilibration time from months to approximately 15 days [7]
- Direct measurement: Considered reference method but suffers from analytical challenges for highly hydrophobic compounds with detection limit issues [7]
- Cosolvent approach: Enables measurement of challenging compounds but may introduce nonlinear behavior affecting extrapolation accuracy [7]
Computational Models:
- LSER models: Demonstrate exceptional predictive accuracy with R² = 0.991 and RMSE = 0.264 for calibration, and R² = 0.985 with RMSE = 0.352 for external validation using experimental descriptors [5] [10]
- Log-linear models: Provide reasonable estimates for nonpolar compounds (R² = 0.985, RMSE = 0.313) but perform poorly for polar compounds (R² = 0.930, RMSE = 0.742) [5]
- QSPR packages: Show varying uncertainty characteristics, with IFSQSAR providing native prediction intervals that capture 90% of external validation data, while OPERA and EPI Suite require adjustment factors of 4 and 2 respectively for similar coverage [17]
Applicability Domain Assessment
Each methodology exhibits distinct strengths and limitations across chemical space:
- LSER models: Cover broad chemical diversity (MW: 32-722, log Ki,O/W: -0.72 to 8.61) with consistent performance [5]
- Three-phase experimental method: Particularly advantageous for highly hydrophobic compounds that challenge conventional methods [7]
- QSPR approaches: Struggle with specific chemical classes including PFAS, ionizable organic chemicals (particularly strong acids and bases), and complex multifunctional structures [17]
- Log-linear correlations: Limited to nonpolar compounds with low hydrogen-bonding capability [5]
Table 4: Integrated Method Selection Guide Based on Research Objectives
| Research Context | Recommended Primary Method | Complementary Approaches | Key Considerations |
|---|---|---|---|
| High-throughput screening | QSPR models (IFSQSAR/OPERA) with uncertainty assessment | Limited experimental validation for critical compounds | Verify applicability domain; assess uncertainty [17] |
| Regulatory submission | Experimental determination (three-phase for HOCs, conventional for others) | LSER prediction for context and justification | Prefer methods with demonstrated accuracy; document comprehensively [7] [5] |
| Research method development | Three-phase system for expanded compound range | Comparison with established literature values | Focus on compounds with literature data for benchmarking [7] |
| Polar compound assessment | LSER models with experimental descriptors | Direct measurement with purified LDPE | Note polarity-dependent differences between pristine and purified LDPE [5] |
Equilibrium partitioning between LDPE and water fundamentally governs leachable accumulation when equilibrium establishes within a product's shelf-life. Contemporary methodologies for characterizing these partition relationships offer complementary strengths—from the experimental robustness of the three-phase system that expands measurable compound range while reducing equilibration time, to the predictive power of LSER models that deliver exceptional accuracy across broad chemical space. The validation of LSER predictions against comprehensive experimental datasets provides researchers with verified computational tools that complement traditional experimental approaches. An integrated strategy—combining in silico screening with targeted experimental verification—represents the most efficient paradigm for comprehensive leachable risk assessment. The methodological comparisons and performance benchmarks presented herein provide drug development professionals with evidence-based guidance for selecting appropriate methodologies based on specific research objectives, compound characteristics, and regulatory requirements.
Implementing LSER Models: From Theory to Practical Prediction
For researchers and drug development professionals, accurately predicting the partitioning of chemical compounds between plastic materials and aqueous phases is a critical aspect of risk assessment for pharmaceutical packaging. The accumulation of leachables in clinically relevant media is principally driven by the equilibrium partition coefficient between the polymer and the medium phase [10]. When leaching equilibrium is reached within a product's duty cycle, these partition coefficients dictate the maximum accumulation of a leachable and thus potential patient exposure [4]. The Linear Solvation Energy Relationship (LSER) model represents a high-performing, mechanistically grounded approach for predicting these partition coefficients, moving beyond coarse estimations that have traditionally limited exposure assessments [6] [4].
This guide provides a comprehensive framework for calculating the partition coefficient between low-density polyethylene (LDPE) and water (log K_i,LDPE/W) using the specifically calibrated LSER equation. The model was developed using experimental data for 159 chemically diverse compounds, spanning a wide range of molecular weight, vapor pressure, aqueous solubility, and polarity, making it particularly suitable for predicting the behavior of potential leachables from pharmaceutical plastics [4].
The LSER Equation for LDPE/Water Partitioning
Core LSER Equation
The foundational LSER model for partitioning between LDPE and water, as calibrated in Part I of the seminal study by Egert et al., is expressed as follows [10] [6] [4]:
log K_i,LDPE/W = -0.529 + 1.098E - 1.557S - 2.991A - 4.617B + 3.886V
This equation enables the prediction of the partition coefficient for any neutral compound with known LSER solute descriptors.
Solute Descriptors and Their Physical Meaning
The LSER model utilizes five key solute descriptors that capture distinct aspects of molecular interaction potential. The table below details each descriptor's significance and interpretation:
Table 1: LSER Solute Descriptors and Their Significance
| Descriptor | Symbol | Interaction Type Represented | Interpretation |
|---|---|---|---|
| Excess Molar Refraction | E | Solute's ability to participate in polarizability interactions via π- and n-electrons | Higher values indicate greater polarizability |
| Dipolarity/Polarizability | S | Solute's combined dipolarity and polarizability | Higher values indicate stronger dipole moments and polarizability |
| Hydrogen Bond Acidity | A | Solute's ability to donate a hydrogen bond | Higher values indicate stronger hydrogen bond donor capability |
| Hydrogen Bond Basicity | B | Solute's ability to accept a hydrogen bond | Higher values indicate stronger hydrogen bond acceptor capability |
| McGowan's Characteristic Volume | V | Solute's molecular size, related to dispersion interactions and cavity formation | Higher values indicate larger molecular volume |
The coefficients in the LSER equation (e.g., -1.557 for S, -4.617 for B) are the system parameters specific to the LDPE/water system. These represent the complementary properties of the phases and determine how sensitive the partitioning is to each solute characteristic [1]. The large negative coefficients for A and B indicate that hydrogen-bonding interactions strongly disfavor partitioning from water into the hydrophobic LDPE polymer, as creating cavity in water for a hydrogen-bonding molecule is energetically costly [10].
Step-by-Step Calculation Procedure
Obtaining the Necessary Solute Descriptors
The first critical step is acquiring the five LSER solute descriptors (E, S, A, B, V) for the compound of interest. Researchers have two primary pathways for this, each with distinct advantages.
Figure 1: Workflow for Calculating log K_i,LDPE/W
Option 1: Experimental Solute Descriptors (Recommended)
The most accurate approach utilizes experimentally determined LSER solute descriptors [10]. These can be retrieved from curated databases or the primary scientific literature. The use of experimental descriptors was validated in Part II of the study, where calculation for an independent validation set (n=52) yielded excellent statistics: R² = 0.985 and RMSE = 0.352 [10]. This represents the highest level of predictive accuracy achievable with the model.
Option 2: Predicted Solute Descriptors
When experimental descriptors are unavailable, researchers can use solute descriptors predicted from the compound's chemical structure using a Quantitative Structure-Property Relationship (QSPR) prediction tool [10] [1]. This approach is more accessible but comes with a slight reduction in predictive accuracy. The validation study using predicted descriptors yielded R² = 0.984 and RMSE = 0.511 [10]. This level of performance is considered indicative for extractables with no experimental descriptors available and remains sufficient for many application purposes.
Performing the Calculation
Once the solute descriptors are obtained, the calculation proceeds through direct substitution into the core LSER equation:
- Substitute the values for E, S, A, B, and V into the equation:
log K_i,LDPE/W = -0.529 + 1.098*E - 1.557*S - 2.991*A - 4.617*B + 3.886*V - Perform the arithmetic operations to compute the value of log K_i,LDPE/W.
- Interpret the result: A higher positive value indicates a stronger tendency for the solute to partition into the LDPE phase rather than the aqueous phase.
Worked Calculation Example
Consider a hypothetical compound with the following solute descriptors:
- E = 0.75
- S = 0.50
- A = 0.10
- B = 0.20
- V = 1.20
The calculation would proceed as follows:
log K_i,LDPE/W = -0.529 + 1.098*(0.75) - 1.557*(0.50) - 2.991*(0.10) - 4.617*(0.20) + 3.886*(1.20)
Breaking down the components:
- Constant: -0.529
- E term: 1.098 * 0.75 = 0.8235
- S term: -1.557 * 0.50 = -0.7785
- A term: -2.991 * 0.10 = -0.2991
- B term: -4.617 * 0.20 = -0.9234
- V term: 3.886 * 1.20 = 4.6632
Summing the components:
log K_i,LDPE/W = -0.529 + 0.8235 - 0.7785 - 0.2991 - 0.9234 + 4.6632 = 2.9567
This result (approximately 2.96) suggests the compound has a moderate to strong preference for the LDPE phase over water.
Experimental Validation and Benchmarking
Model Performance Metrics
The LSER model for LDPE/water partitioning has undergone rigorous validation. The following table summarizes its performance characteristics based on the comprehensive study by Egert et al.:
Table 2: LSER Model Performance for log K_i,LDPE/W Prediction
| Performance Aspect | Calibration (Part I) | Validation with Experimental Descriptors | Validation with QSPR Descriptors |
|---|---|---|---|
| Dataset Size (n) | 156 | 52 | 52 |
| Coefficient of Determination (R²) | 0.991 | 0.985 | 0.984 |
| Root Mean Square Error (RMSE) | 0.264 | 0.352 | 0.511 |
| Molecular Weight Range | 32 to 722 | Not specified | Not specified |
| log K_i,LDPE/W Range | -3.35 to 8.36 | Not specified | Not specified |
The high R² values across all scenarios demonstrate the model's strong explanatory power, while the low RMSE values indicate high predictive precision [10] [4]. The slight increase in RMSE when using predicted descriptors reflects the additional uncertainty introduced by the descriptor prediction step.
Comparison with Alternative Prediction Methods
Log-Linear Octanol-Water Model
A common alternative approach involves using a log-linear correlation with the octanol-water partition coefficient (log K_i,O/W). The performance of this simpler model is highly dependent on the chemical nature of the solute [4]:
- For nonpolar compounds (low hydrogen-bonding propensity):
log K_i,LDPE/W = 1.18 log K_i,O/W - 1.33(n=115, R²=0.985, RMSE=0.313) - For polar compounds (including mono-/bipolar compounds): The correlation weakens significantly (n=156, R²=0.930, RMSE=0.742)
The LSER model's superiority is particularly evident for polar compounds, where it accounts for specific hydrogen-bonding and polar interactions that the log-linear model cannot capture effectively [4].
Cosolvency Models for Water-Ethanol Mixtures
In pharmaceutical applications, water-ethanol mixtures are frequently used as simulating solvents for leaching studies. Research has evaluated the ability of cosolvency models, combined with the LSER approach, to predict solute partitioning between LDPE and water-ethanol mixtures [14]. The LSER-based cosolvency model was found to be slightly superior to the log-linear cosolvency model (Yalkowsky's model), with both enabling the tailored preparation of simulating solvent mixtures that reflect the extraction strength of clinically relevant media [14].
Advanced Applications and Research Implications
Comparison with Other Polymers
The LSER framework allows for direct comparison of LDPE's sorption behavior with other polymers used in pharmaceutical applications. Based on LSER system parameters, LDPE can be compared to polydimethylsiloxane (PDMS), polyacrylate (PA), and polyoxymethylene (POM) [10]:
- PA and POM, with their heteroatomic building blocks, exhibit stronger sorption for polar, non-hydrophobic compounds in the log K_i,LDPE/W range of 3 to 4, due to their capabilities for polar interactions.
- For compounds with log K_i,LDPE/W > 4, all four polymers exhibit roughly similar sorption behavior, as hydrophobic interactions become dominant.
Amorphous Polymer Partitioning
For more theoretical comparisons with liquid phases, partition coefficients can be converted to represent partitioning into the amorphous fraction of LDPE only (log Ki,LDPEamorph/W). A LSER model recalibrated on this basis changes the constant in the equation from -0.529 to -0.079, making it more similar to a corresponding LSER model for n-hexadecane/water systems [10]. This adjustment can be valuable when comparing polymer partitioning directly with partitioning into liquid organic phases.
Researcher's Toolkit
Table 3: Essential Research Reagents and Resources for LSER Applications
| Resource Category | Specific Examples/Functions | Application in LSER Studies |
|---|---|---|
| Polymer Materials | Purified Low-Density Polyethylene (LDPE) | Sorption studies; determining experimental partition coefficients [4] |
| Chemical Standards | Chemically diverse compounds with known descriptors | Model calibration and validation [10] [4] |
| LSER Database | Curated database of solute descriptors (E, S, A, B, V) | Input parameters for partition coefficient calculations [10] [1] |
| QSPR Prediction Tools | Software for predicting LSER descriptors from structure | Generating descriptors when experimental values are unavailable [10] |
| Simulating Solvents | Water-ethanol binary mixtures | Modeling leaching into clinically relevant media [14] |
The LSER approach for predicting log K_i,LDPE/W represents a robust, accurate, and mechanistically grounded methodology that significantly advances the predictive modeling capabilities available to pharmaceutical researchers and regulatory scientists. With its high precision (R² > 0.99, RMSE ~0.26-0.35) and applicability across a broad chemical space, it enables more reliable estimation of leachable accumulation and patient exposure from plastic packaging and medical devices [10] [4].
The step-by-step guide presented here, from descriptor acquisition to final calculation, provides researchers with a practical framework for implementing this powerful model. The benchmarking data confirms its superiority over simpler log-linear models, particularly for polar compounds, while the ongoing development of interconnected thermodynamic frameworks, such as Partial Solvation Parameters (PSP), promises to further enhance the extraction and utilization of thermodynamic information from the LSER database [1]. As the field progresses, the integration of LSER-predicted partition coefficients with cosolvency models and thermodynamic cycles will continue to increase the reliability of chemical safety risk assessments while optimizing resource allocation in pharmaceutical development.
Sourcing Experimental vs. QSPR-Predicted Solute Descriptors
In the field of environmental chemistry and pharmaceutical sciences, predicting the partitioning behavior of organic compounds between polymers, such as low-density polyethylene (LDPE), and water is crucial for assessing environmental transport, fate, and human exposure to leachables. Linear Solvation Energy Relationships (LSERs) are a powerful predictive tool in this domain. The accuracy of these models is fundamentally tied to the quality of the solute descriptors used as inputs. These descriptors, which encode key molecular properties, can be sourced either through experimental determination or via Quantitative Structure-Property Relationship (QSPR) predictions. This guide provides an objective comparison of these two sourcing methodologies, offering a structured framework for researchers to select the appropriate approach based on their project's requirements for accuracy, resources, and applicability.
Performance Comparison: Experimental vs. QSPR-Predicted Descriptors
The choice between experimental and QSPR-predicted solute descriptors has a direct and quantifiable impact on the predictive performance of LSER models. The following table summarizes the key performance indicators derived from a robust validation study on an LSER model for LDPE/water partition coefficients (log K_{i, LDPE/W}) [6] [10].
Table 1: Performance Benchmark of an LDPE/Water LSER Model Using Different Solute Descriptor Sources
| Descriptor Source | Number of Compounds (n) | Coefficient of Determination (R²) | Root Mean Square Error (RMSE) | Key Characteristics |
|---|---|---|---|---|
| Experimental Solute Descriptors | 52 | 0.985 | 0.352 | Considered the benchmark for accuracy and precision; requires reference data or laborious experimental work. |
| QSPR-Predicted Solute Descriptors | 52 | 0.984 | 0.511 | Provides a practical alternative for compounds with no experimental data; introduces higher uncertainty. |
Interpretation of Comparative Data
- Accuracy and Precision: While both descriptor sources yield a high R² value (~0.984), indicating a strong model structure, the ~45% higher RMSE associated with QSPR-predicted descriptors is significant. This increase in error is attributed to the inherent uncertainty introduced when predicting the solute descriptors solely from a compound's chemical structure [6]. The model based on experimental descriptors (RMSE = 0.352) is therefore considered more accurate and precise [10].
- Application Domain: The performance of QSPR-predicted descriptors is highly indicative for "extractables" with no experimental descriptors available [6]. Their use is excellent for high-throughput screening and initial risk assessments. However, for definitive safety evaluations or regulatory submissions where higher accuracy is critical, experimentally-derived descriptors are the superior choice.
Experimental Protocols for Descriptor Determination and Model Validation
Determining Solute Descriptors from Chromatographic Measurements
A foundational method for obtaining experimental solute descriptors involves chromatographic techniques [18]. The general workflow involves:
- Measurement: Gas-liquid chromatographic (GLC) retention data or reverse-phase high-performance liquid chromatographic (RP-HPLC) measurements are collected for the solute of interest across multiple systems [18].
- Regression Analysis: The retention data are processed through multiple linear regression against a system of equations that describe the solvation properties of the chromatographic phases.
- Descriptor Extraction: This analysis yields a set of solute descriptors (e.g., E, S, A, B, V, L) that encode the molecule's excess molar refractivity, dipolarity/polarizability, hydrogen-bond acidity/basicity, and molecular volume [19] [18]. These experimentally-determined descriptors are considered highly reliable.
Validating QSPR Models with Experimental Partitioning Data
To build confidence in QSPR-based predictions, rigorous experimental validation is essential. A study on polychlorinated biphenyls (PCBs) exemplifies a robust protocol for validating K_{PE-w} predictions [20]:
- Model Development: Several QSPR models (e.g., using Random Forest, Support Vector Machine) are developed to predict
log K_{PE-w}based on molecular descriptors. - Experimental Verification: A three-phase system (aqueous phase, surfactant micelles, LDPE) is used to experimentally determine the
log K_{PE-w}values for 16 PCBs. This system accelerates equilibration time compared to conventional two-phase methods [20]. - Performance Assessment: The modeled results are compared against the experimental values. The study reported strong agreement, with residuals within ± 0.3 log units, thereby validating the predictive capability of the QSPR models [20].
The Scientist's Toolkit: Key Research Reagents and Materials
Table 2: Essential Materials for LDPE-Water Partitioning Studies
| Material/Reagent | Function in Research |
|---|---|
| Low-Density Polyethylene (LDPE) | The polymeric phase of interest; used in passive sampling devices (PSDs) to study the partitioning and environmental fate of contaminants [6] [20]. |
| Aqueous Buffers | Represents the aqueous phase (e.g., environmental water, clinically relevant media); used to establish partitioning equilibrium under controlled pH conditions [4]. |
| Organic Solvents (e.g., Methanol) | Used as co-solvents in experimental methods to increase equilibration rates and stabilize solutions for highly hydrophobic compounds [20]. |
| Surfactants | Used to form micelles in three-phase (LDPE/micelle/water) systems, which drastically reduce equilibration time for experimental validation of partition coefficients [20]. |
| Reference Compounds | Compounds with well-established experimental solute descriptors and partition coefficients; crucial for calibrating instruments and validating LSER model predictions [6] [18]. |
| Polychlorinated Biphenyls (PCBs) | Commonly studied hydrophobic organic contaminants (HOCs); serve as model solutes in developing and validating QSPR models for polymer-water partitioning [20]. |
The accurate prediction of chemical migration is a critical component in the safety evaluation of medical devices. Within a regulatory framework governed by standards like ISO 10993-18, demonstrating biocompatibility often hinges on understanding which chemical substances may be released from a device material into a patient's body [21]. For devices incorporating low-density polyethylene (LDPE) or similar polymers, the partition coefficient between the polymer and water (Kpe-w) is a key parameter for estimating patient exposure to potential leachables.
This case study is situated within a broader thesis validating the use of Linear Solvation Energy Relationships (LSERs) for predicting LDPE/water partition coefficients. Traditionally, obtaining these coefficients relies on laborious and lengthy experimental methods, which are particularly challenging for highly hydrophobic compounds that can take weeks or even a year to reach equilibrium [7]. Computational models like LSERs offer a powerful alternative, enabling robust and accurate predictions of partitioning behavior. This study objectively compares the traditional experimental approach with the emerging methodology of LSER prediction, providing experimental data and protocols to guide researchers in designing more efficient and informative extraction experiments for medical devices.
Background: Extraction Testing for Medical Devices
Extraction testing is a foundational practice in the biological safety assessment of medical devices. As outlined in ISO 10993-12, its purpose is to obtain a sample that can be used for either biological testing or chemical analysis to support a toxicological risk assessment [21] [22]. The goal is to simulate, or deliberately exceed, what the human body might extract from a device during clinical use.
Extraction Study Approaches
The ISO 10993-18 standard delineates three primary extraction approaches, selected based on the device's contact duration and nature [21]:
- Simulated-Use Extraction: Aims to estimate the release of substances under actual clinical conditions of use.
- Exaggerated Extraction: Uses conditions (e.g., higher temperature, longer time) that are more severe than clinical use to increase the number and amount of chemical constituents extracted. The U.S. Food and Drug Administration (FDA) often recommends this approach for devices with limited exposure duration [21].
- Exhaustive Extraction: An iterative series of extractions performed until the amount of material extracted in a subsequent step is less than 10% of the initial extraction. This is strongly encouraged for devices with long-term exposure (exceeding 30 days) [21].
The selection of an extraction strategy is a critical first step in device evaluation, directly influencing the experimental design and the relevance of the resulting data for safety assessments.
Comparative Methodologies: Experimental vs. LSER Prediction
This section details the two core methodologies for determining Kpe-w, a vital parameter for translating measured concentrations in a polymer into estimated patient exposure.
Traditional Experimental Method
The conventional method involves a direct two-phase system where the polymer is immersed in water until equilibrium is reached.
Experimental Protocol
- Materials: LDPE sheets (often purified via solvent extraction to remove interferents), high-purity water, hydrophobic organic compounds (HOCs) of interest (e.g., PAHs, PCBs, PBDEs) [7] [5].
- Procedure:
- LDPE films are cut to a specific surface area and added to aqueous solutions containing the target HOCs.
- The system is agitated and maintained at a constant temperature (e.g., 25°C) for a duration sufficient to reach equilibrium. For very hydrophobic compounds, this can be extremely long, up to 365 days [7].
- Upon equilibrium, the concentration of the chemical in both the polymer phase (
Cpe) and the water phase (Cw) is measured. - The partition coefficient is calculated as
Kpe-w = Cpe / Cw.
Limitations and Innovations
A major limitation is the long equilibration time and the analytical difficulty of measuring trace concentrations in water. A novel three-phase system has been developed to address these issues [7]. This method introduces a surfactant (e.g., Brij 30) to create a micellar pseudo-phase. The Kpe-w is then determined from a combination of two more easily measured values: the micelle-water partition coefficient (Kmic-w) and the LDPE-micelle partition coefficient (Kpe-mic). This approach can shorten equilibration time to approximately half a month and avoids the analytical challenges of direct aqueous phase concentration measurement [7].
LSER Prediction Method
Linear Solvation Energy Relationships (LSERs) are poly-parameter models that predict partition coefficients based on a compound's molecular descriptors, accounting for the various intermolecular interactions involved in the partitioning process [23] [5].
Model Calibration and Equation
A robust LSER model for the LDPE-water system was recently calibrated using a diverse set of 159 compounds. The model is defined by the following equation [5]:
log Ki,LDPE/W = -0.529 + 1.098E - 1.557S - 2.991A - 4.617B + 3.886V
Table 1: Abraham Solute Parameters in the LSER Model
| Symbol | Molecular Interaction Descriptor | Interpretation in LDPE-Water Partitioning |
|---|---|---|
| E | Excess molar refractivity | Polarizability/Van der Waals interactions; favors LDPE phase |
| S | Dipolarity/polarizability | Dipolar interactions; strongly favors water phase |
| A | Hydrogen-bond acidity | Hydrogen-bond donating ability; strongly favors water phase |
| B | Hydrogen-bond basicity | Hydrogen-bond accepting ability; strongly favors water phase |
| V | McGowan molecular volume | Dispersion interactions; strongly favors LDPE phase |
This model demonstrates high accuracy and precision (n = 156, R² = 0.991, RMSE = 0.264) across a wide chemical space [5].
Advantages Over One-Parameter Models
While simpler one-parameter models based on the octanol-water partition coefficient (log Kow) exist, they are less accurate, especially for polar compounds. The LSER model's superiority stems from its ability to account for all relevant intermolecular interactions, not just hydrophobicity [23] [5]. For nonpolar compounds, a log Kow-based model (log Ki,LDPE/W = 1.18 log Ki,O/W - 1.33) can be useful, but its performance degrades significantly when applied to mono- or bipolar compounds [5].
Comparative Performance Data
The following tables summarize the quantitative performance of the different methodologies and models discussed.
Table 2: Performance Comparison of Kpe-w Prediction Models
| Model Type | Representative Equation | Applicability | Accuracy (RMSE) | Key Advantage |
|---|---|---|---|---|
| LSER (pp-LFER) | log K = -0.529 + 1.098E - 1.557S - 2.991A - 4.617B + 3.886V |
Diverse chemicals, including polar compounds [5] | 0.264 [5] | Accounts for all major molecular interactions |
| Octanol-Water (op-LFER) | log Ki,LDPE/W = 1.18 log Ki,O/W - 1.33 |
Nonpolar compounds with low H-bonding propensity [5] | 0.313 (nonpolar); 0.742 (full set) [5] | Simple, uses a widely available parameter |
| Hexadecane-Water (op-LFER) | Linear regression vs. log Khexadecane-w [23] |
Better proxy for LDPE than octanol [23] | ~0.41-0.42 [23] | More accurately represents Van der Waals interactions |
Table 3: Comparison of Experimental Methodologies for Determining Kpe-w
| Experimental Method | Typical Equilibration Time | Key Challenge | Best For |
|---|---|---|---|
| Traditional Two-Phase | Up to 365 days for very hydrophobic compounds [7] | Measuring very low aqueous concentrations (Cw); long duration [7] | Fundamental research; model validation |
| Novel Three-Phase (Micellar) | ~15 days [7] | Requires measurement of two partition coefficients [7] | Rapid generation of high-quality experimental data for model training |
Application in Medical Device Experiment Design
The integration of LSER predictions and modern experimental methods provides a powerful toolkit for designing efficient and representative extraction experiments for medical devices.
A Workflow for Informed Extraction Design
The following diagram illustrates a strategic workflow for applying LSERs and modeling insights to the design of medical device extraction studies.
Informed Extraction Experiment Design Workflow
Integrating LSERs with Extraction Models
Physics-based mass transport models can predict extraction efficiency (M/M₀, the fraction of leachable mass released) and extract concentration (C/C₀) based on dimensionless parameters [24]:
- Ψ (Thermodynamic parameter):
Ψ = Vs / (Vp * K), whereVsis solvent volume,Vpis polymer volume, andKis the partition coefficient. - τ (Kinetic parameter):
τ = Dt / L², whereDis the diffusion coefficient,tis time, andLis a characteristic length (e.g., thickness).
LSER-predicted Kpe-w values can be directly input as K into these models to simulate and optimize extraction conditions in silico before lab work begins. This allows researchers to answer critical design questions, such as whether an exhaustive extraction is achievable or how the surface-area-to-volume ratio will impact the concentration used in biological testing [24]. The model reveals that complete extraction (M/M₀ approaches 1) only occurs for relatively large values of Ψ (≥10) and τ (≥1) [24].
The Scientist's Toolkit: Key Research Reagents and Materials
Table 4: Essential Materials for LDPE/Water Partitioning Research
| Item | Function/Application | Notes |
|---|---|---|
| Purified LDPE | Polymer phase for partitioning experiments | Purification via solvent extraction removes interferents, which is critical for accurate sorption measurements [5]. |
| Hydrophobic Organic Compounds (HOCs) | Model leachables for method development/validation | Include PAHs, PCBs, PBDEs to cover a range of hydrophobicity and polarities [7] [5]. |
| Surfactant (e.g., Brij 30) | Forms micellar pseudo-phase in the three-phase experimental system | Enhances solute solubility, drastically reducing equilibration time [7]. |
| Aqueous Buffers | Aqueous phase for partitioning | Controls pH to simulate physiological conditions or standardize tests [5]. |
| Abraham Solute Parameters (E, S, A, B, V) | Descriptors for LSER predictions | Found in chemical databases or calculated using specialized software [23] [5]. |
This case study demonstrates a paradigm shift in designing extraction experiments for medical devices. While traditional experimental methods remain the foundational source of truth, they are often slow and resource-intensive. The emergence of robust and highly accurate LSER models provides researchers with a powerful predictive tool to streamline the design process.
The strategic approach involves using LSERs to predict critical partition coefficients and then leveraging physics-based models to simulate extraction scenarios. This integrated methodology enables a more principled selection of extraction conditions—such as solvent volume, time, and temperature—that are truly worst-case and scientifically justified. By reducing reliance on trial-and-error in the laboratory, researchers can develop more efficient and effective extraction protocols, ultimately accelerating the safety evaluation of medical devices while ensuring the highest level of patient protection. The validation of these computational tools, as part of a broader thesis, marks a significant advancement in the field of medical device biocompatibility assessment.
In the fields of medical device safety and drug development, accurately predicting the distribution of chemicals from materials into the body is paramount for toxicological risk assessment. While substantial research has focused on predicting polymer-water partitioning, this represents only the first step in understanding potential patient exposure. The critical subsequent step involves forecasting how these leached compounds distribute within the body, particularly to blood and adipose tissue, which govern systemic circulation and long-term accumulation, respectively. Traditional experimental approaches for determining these partition coefficients are resource-intensive and low-throughput, creating a critical need for robust predictive models. Among the various computational approaches, Linear Solvation Energy Relationship (LSER) models have emerged as powerful tools for translating polymer-water partitioning data into biologically relevant distribution predictions. This guide objectively compares the performance of LSER methodologies against conventional surrogate solvent systems, providing researchers with validated protocols and data-driven insights for implementing these approaches in safety assessment workflows.
Table 1: Key Partition Coefficients in Medical Product Safety Assessment
| Partition Coefficient | Symbol | Role in Risk Assessment |
|---|---|---|
| Low-Density Polyethylene/Water | KLDPE/W | Predicts leaching potential from a common polymer material into aqueous solutions. |
| Blood/Water | KBlood/W | Estimates solubility and distribution in systemic circulation. |
| Adipose Tissue/Water | KAT/W | Predicts long-term storage and accumulation in body fat. |
| Adipose Tissue/Plasma | KAT/P | Describes tissue distribution of drugs at steady state. |
Comparative Performance of Predictive Methodologies
LSER Models for Blood and Adipose Tissue Partitioning
The application of LSER models represents a significant advancement in predicting tissue partitioning. A 2023 study developed LSER models specifically to predict partitioning from low-density polyethylene (LDPE) to blood and adipose tissue, with the goal of optimizing the design of extraction experiments for medical device safety assessment [25]. When benchmarked against traditional solvent surrogates, the performance of this LSER approach was notable. For predicting blood/water partition coefficients, the LSER model performed equally well as a 60:40 ethanol/water mixture and outperformed surrogate solvents like octanol and butanol, as measured by root mean squared error (RMSE) [25]. For adipose tissue/water partitioning, the experimentally determined octanol/water partition coefficient (KOW) showed the best performance, but the RMSE of the LSER approach was in the same range, demonstrating its competitive predictive capability [25]. This indicates that LSER models provide a reliable, single-method framework for predicting partitioning to both blood and adipose tissue, reducing the need for multiple surrogate systems.
Tissue Composition-Based Models and Surrogate Solvents
Beyond LSERs, tissue composition-based models provide another important predictive approach. These models use fundamental physiological data—such as the lipid and water content of tissues and blood—in combination with compound-specific parameters to estimate partition coefficients. A critical finding from this research area is that olive oil is a substantially better surrogate for adipose tissue lipids than n-octanol [26] [27]. One seminal study demonstrated that predictions of in vivo adipose tissue:plasma partition coefficients (KAT/P) using olive oil-water data differed from experimental values by an average factor of only 1.17, whereas predictions using n-octanol-water data differed by factors of 15.0 and 40.7, depending on the calculation method [26] [27]. Furthermore, for highly lipophilic organic compounds (HLOCs with log KOW > 4), the partition coefficient between adipose tissue and blood (KAT/B) becomes largely independent of the specific chemical's identity. Instead, it plateaus at a value approximately equal to the ratio of the lipid content in adipose tissue to that in blood, as the chemicals reside almost exclusively in the lipid fractions of both compartments [28].
Table 2: Performance Comparison of Predictive Methods for Tissue Partitioning
| Prediction Method | Target Tissue | Key Input Parameters | Reported Performance Metric | Advantages |
|---|---|---|---|---|
| LSER Model [25] | Blood & Adipose Tissue | LSER solute descriptors (E, S, A, B, V) | RMSE competitive with best surrogates | Single unified model for multiple tissues; accounts for multiple molecular interactions |
| Octanol/Water (KOW) [25] [26] | Adipose Tissue | n-octanol:water partition coefficient | Best performer for adipose tissue (RMSE); but poor for KAT/P prediction (avg. factor error 15-40x) [26] | Simple, widely available data; good for very lipophilic compounds |
| Olive Oil/Water [26] [27] | Adipose Tissue | Olive oil:water partition coefficient | Excellent for KAT/P (avg. factor error 1.17x) [26] | Superior lipid surrogate; accounts for triglyceride compatibility |
| Ethanol/Water (60:40) [25] | Blood | Solvent composition | RMSE similar to LSER | Good blood surrogate for extraction studies |
Experimental Protocols for Key Methodologies
LSER Model Development and Validation Protocol
The robust LSER models cited in this guide were built upon a rigorous experimental foundation. The core dataset was developed by determining partition coefficients between low density polyethylene (LDPE) and aqueous buffers for 159 compounds spanning an extensive range of chemical diversity, molecular weight (32 to 722 Da), and hydrophobicity (log KLDPE/W from -3.35 to 8.36) [4]. This chemical space is considered representative of the universe of compounds potentially leaching from plastics. The resultant LSER model for LDPE-water partitioning takes the form [6] [10] [4]:
log Ki,LDPE/W = -0.529 + 1.098E - 1.557S - 2.991A - 4.617B + 3.886V
The model's accuracy and precision were rigorously validated (n = 156, R² = 0.991, RMSE = 0.264) [4]. A critical validation step involved setting aside ~33% of the data (n = 52) as an independent validation set. When predictions for this set were made using experimental LSER solute descriptors, they yielded R² = 0.985 and RMSE = 0.352 [6] [10]. Even when using predicted descriptors from a QSPR tool, the model maintained strong performance (R² = 0.984, RMSE = 0.511), confirming its utility for compounds without experimentally determined descriptors [6] [10].
Three-Phase Partitioning Measurement Protocol
To address challenges in measuring LDPE-water partition coefficients for highly hydrophobic organic compounds (HOCs), a novel three-phase partitioning system has been developed [7]. This method significantly improves upon traditional two-phase systems by reducing equilibration time and minimizing analytical errors associated with measuring low aqueous concentrations. The procedural steps are as follows:
- System Setup: A surfactant (Brij 30) is added to a traditional LDPE-water system at a concentration above its critical micelle concentration (CMC) to form a micellar pseudo-phase.
- Measurement of Kmic-w: The micelle-water partition coefficient (Kmic-w) is determined by measuring the enhancement of chemical solubility in Brij 30 solutions above the CMC. The relationship between total solubility and surfactant concentration is linear, with the slope representing Kmic-w [7].
- Measurement of KPE-mic: The LDPE-micelle partition coefficient (KPE-mic) is directly measured as the equilibrium ratio of solute concentration in the LDPE phase to its concentration in the micellar phase. Concentrations in both organic phases are sufficiently high to avoid analytical detection issues [7].
- Calculation of KPE-w: The desired LDPE-water partition coefficient is calculated from the two experimentally measured values: KPE-w = KPE-mic × Kmic-w [7].
This method reduces equilibration time to approximately half a month, compared to much longer periods (e.g., up to 365 days for some PBDEs) in direct measurement methods, while providing high accuracy [7].
Essential Research Reagent Solutions
Successful implementation of the described predictive models and experimental protocols requires specific research reagents and computational tools. The following table details key solutions and their functions in partitioning research.
Table 3: Essential Research Reagent Solutions for Partitioning Studies
| Reagent / Tool | Function in Research | Application Context |
|---|---|---|
| Low-Density Polyethylene (LDPE) | Model polymer phase for partitioning experiments; common medical device material. | Experimental determination of KLDPE/W [6] [7] [4] |
| Brij 30 Surfactant | Forms micellar pseudo-phase in three-phase partitioning systems. | Enables accurate measurement of KPE-w for HOCs [7] |
| n-Octanol | Surrogate solvent for lipophilic phases in traditional partition models. | Determination of KOW, a common lipophilicity descriptor [25] [26] |
| Olive Oil | Physiologically relevant surrogate for adipose tissue lipids. | Prediction of adipose tissue:plasma partition coefficients [26] [27] |
| Ethanol/Water Mixtures | Simulating solvents for blood and other biological fluids. | Extraction studies for medical device leachables [25] |
| LSER Solute Descriptors | Compound-specific parameters describing molecular interactions. | Core input variables for LSER model predictions [25] [6] |
The comparative data and protocols presented in this guide provide a evidence-based foundation for selecting appropriate methodologies to predict chemical partitioning to blood and adipose tissue. The LSER approach offers a unified, robust framework that performs competitively with the best surrogate systems for blood and provides respectable accuracy for adipose tissue, making it an excellent choice for a standardized in-silico tool within a risk assessment workflow [25]. For applications specifically focused on adipose tissue distribution, especially for ionizable drugs, tissue composition-based models using olive oil as a lipophilicity surrogate remain the gold standard due to their superior accuracy [26] [27]. The choice between these methods should be guided by the specific context: LSER models are particularly powerful for integrating material leaching and biological distribution into a single predictive framework, while olive oil-based models are ideal for detailed pharmacokinetic modeling of tissue distribution. The ongoing refinement of these models, coupled with innovative experimental methods like three-phase partitioning, continues to enhance our ability to accurately forecast chemical behavior in biological systems, ultimately supporting the development of safer medical products and more effective therapeutics.
Tailoring Simulating Solvents (e.g., Ethanol/Water Mixtures) Using LSER Projections
In the pharmaceutical and medical device industries, evaluating the safety of primary contact plastic materials is paramount. Such evaluations require extractables and leachables studies to estimate patient exposure to chemicals that can potentially migrate from the plastic material into the clinically relevant medium (CRM) [29] [30]. However, direct analysis of leachables in the CRM, which can be a complex composition, is often challenging [29] [14]. To overcome this, simulating solvents, notably binary water-ethanol mixtures, are widely used to mimic the polarity-driven extraction strength of the CRM [29] [14] [30].
The central challenge lies in quantitatively tailoring the composition of these simulating solvent mixtures to accurately reflect the extraction propensity of the CRM [29] [30]. This guide compares two key cosolvency models used to predict solute partitioning in these systems, focusing on their application within a framework validated by Linear Solvation Energy Relationship (LSER) projections for Low-Density Polyethylene (LDPE)/water partitioning research.
Model Comparison: Log-Linear vs. LSER-Based Cosolvency
The selection of an appropriate simulating solvent mixture hinges on accurately predicting the partition coefficient of a solute between a polymer (e.g., LDPE) and a water-ethanol mixture. The following table provides a high-level comparison of the two main cosolvency models used for this purpose.
Table 1: Comparison of Cosolvency Models for Predicting Solute Partitioning
| Feature | Log-Linear (LL) Cosolvency Model | LSER-Based Cosolvency Model |
|---|---|---|
| Fundamental Principle | Assumes a log-linear relationship between solubilization power and solute hydrophobicity (often using the octanol/water partition coefficient, log Ki,O/W) [30]. | Based on Abraham-type LSERs, which describe the partition coefficient as a function of multiple solute descriptors capturing different molecular interactions [29] [6] [30]. |
| Theoretical Foundation | Empirical; relatively simple. | Stronger thermodynamical foundation, accounting for specific intermolecular forces [29] [30]. |
| Input Parameters | Cosolvent constants and the solute's log Ki,O/W [30]. | Five experimental LSER solute descriptors (E, S, A, B, V) and system parameters (e, s, a, b, v) [6] [30]. |
| Predictability at Low Ethanol Fractions (e.g., 10-20%) | Good and comparable to the LSER model [30]. | Good and comparable to the log-linear model [30]. |
| Predictability at High Ethanol Fractions (e.g., 35-50%) | Less accurate compared to the LSER model [30]. | Superior; maintains higher accuracy due to its ability to model non-linear solubilization behavior [29] [30]. |
| Key Advantage | Simplicity and ease of use. | Higher accuracy, especially for complex chemical systems and a wider range of solvent compositions. |
Performance Benchmarking and Experimental Validation
Experimental validation across a diverse set of solutes confirms the comparative performance of these models. The following table summarizes quantitative performance data from a study that calculated partition coefficients between LDPE and water-ethanol mixtures at various volume fractions (fEtOH) [30].
Table 2: Experimental Performance Data of Cosolvency Models
| Ethanol Volume Fraction (fEtOH) | Log-Linear Model (Average Absolute Deviation) | LSER-Based Model (Average Absolute Deviation) |
|---|---|---|
| 0.10 | 0.27 | 0.27 |
| 0.20 | 0.29 | 0.28 |
| 0.35 | 0.36 | 0.26 |
| 0.50 | 0.43 | 0.29 |
The data shows that while both models perform similarly at lower ethanol fractions, the LSER-based model demonstrates significantly better predictability at higher ethanol fractions (35% and 50%), with lower average absolute deviations from experimentally obtained values [30]. This is attributed to its more robust foundation in capturing the specific interactions between the solute, water, and ethanol [29].
Experimental Protocols
Core Workflow: From Prediction to Experimental Verification
The general methodology for tailoring simulating solvents and validating the LSER projections involves a combination of predictive modeling and experimental verification. The process can be summarized in the following workflow, which outlines the key stages from initial setup to final model validation.
Detailed Methodologies for Key Experiments
1. Batch Equilibrium Sorption Experiments: This protocol is used to determine the experimental partition coefficient (Ki, LDPE/M) between LDPE and the water-ethanol simulating solvent (medium M) [30].
- LDPE Film Preparation: LDPE films are cut into appropriate sizes (e.g., 2 cm diameter discs). The films are pre-washed and dried to remove any contaminants [30].
- Solute Spiking: The LDPE films are spiked with a carefully selected set of test solutes, typically dissolved in a volatile solvent like methanol. The solvent is allowed to evaporate, leaving the solute on the polymer [30].
- Equilibration: The spiked LDPE films are immersed in the water-ethanol mixture (at specific volume fractions such as 0.1, 0.2, 0.35, and 0.5) in sealed vials. The vials are stored at a controlled temperature (e.g., 40°C) for a defined period (e.g., 21 days) to reach equilibrium, followed by a stabilization period at a lower temperature (e.g., 25°C for 14 days) [30].
- Concentration Analysis: After equilibration, the concentration of the solute in the polymer phase (CP) and in the solvent phase (CM) is quantified using analytical techniques such as Liquid Chromatography with Ultraviolet detection (LC/UV). The partition coefficient is calculated as Ki, LDPE/M = CP / CM [30].
2. LSER Model Application and Prediction:
- Solute Descriptor Acquisition: Experimentally determined Abraham LSER solute descriptors (E, S, A, B, V) for the test compounds are obtained. These descriptors represent excess molar refractivity, dipolarity/polarizability, hydrogen-bond acidity, hydrogen-bond basicity, and McGowan's characteristic volume, respectively [6] [30].
- Partition Coefficient Calculation: The partition coefficient between LDPE and water (log Ki, LDPE/W) is calculated using a previously established and validated LSER model. An example of such a model is [6]:
log K<sub>i, LDPE/W</sub> = -0.529 + 1.098E - 1.557S - 2.991A - 4.617B + 3.886V - Thermodynamic Cycle Application: The hypothetical partition coefficient based on differential solubility between the water-ethanol mixture and pure water, log (Si,fC/Si,W), is calculated using the LSER-based cosolvency model. Finally, the partition coefficient between LDPE and the simulating solvent mixture (log Ki, LDPE/M) is derived by applying a thermodynamic cycle that combines log Ki, LDPE/W and log (Si,fC/Si,W) [29] [14] [30].
The Scientist's Toolkit: Essential Research Reagents and Materials
Successful implementation of this research requires specific materials and an understanding of their function within the experimental and computational framework.
Table 3: Essential Research Reagents and Materials
| Item | Function / Relevance in Research |
|---|---|
| Low-Density Polyethylene (LDPE) Films | The model polymer phase used in sorption experiments. Its properties are well-characterized, making it a standard for studying partitioning behavior in pharmaceutical packaging [29] [30]. |
| Ethanol (Absolute) | The cosolvent used to create binary water-ethanol simulating solvents. It modifies the polarity and solvation strength of the pure aqueous phase [29] [14]. |
| Chemically Diverse Solute Set | A critical set of test compounds with available experimental LSER descriptors. The set should cover a wide range of molecular weights, and hydrogen-bonding properties (acids, bases, neutrals, amphions) to robustly validate the models [6] [30]. |
| Abraham LSER Solute Descriptors (E, S, A, B, V) | Quantitative numerical values that describe a molecule's potential for various intermolecular interactions. These are the essential inputs for the LSER-based cosolvency model [6] [30]. |
| LSER System Parameters (e, s, a, b, v) | Coefficients in the LSER equation that characterize the properties of the specific system being studied (e.g., the LDPE/water-ethanol system). They are derived by correlating experimental partition data with solute descriptors [29] [6]. |
This guide has objectively compared two methodologies for tailoring simulating solvents. The experimental data and model foundations demonstrate that while the log-linear cosolvency model offers simplicity and is sufficient for initial estimates at low cosolvent fractions, the LSER-based cosolvency model provides a more robust and accurate predictive framework. Its superiority is particularly evident at higher ethanol fractions, where it more effectively captures the complex, non-linear solubilization behavior of chemically diverse solutes.
For researchers and drug development professionals, the integration of LSER-predicted partition coefficients with LSER-based cosolvency models represents a powerful strategy. This approach enables the rational, data-driven design of water-ethanol simulating solvents that can more reliably mimic clinically relevant media. The outcome is a significant enhancement in the reliability of patient exposure estimations for leachables, leading to more efficient and scientifically sound chemical safety risk assessments for plastic materials used in pharmaceutical applications [29] [30]. Future work in this field should focus on expanding validation to a broader compound set, exploring other cosolvent systems, and developing integrated software toolkits to streamline this methodology for industrial application.
Enhancing LSER Model Performance and Addressing Limitations
In pharmaceutical development and environmental science, accurately predicting the distribution of chemicals between low-density polyethylene (LDPE) and aqueous phases is critical for assessing patient exposure to leachables and understanding environmental transport of contaminants. Linear Solvation Energy Relationships (LSERs) have emerged as a powerful predictive tool for estimating partition coefficients (log K~i,LDPE/W~), which define the equilibrium distribution of a compound between a polymer and water. These models are particularly valuable for chemical safety risk assessments, where they help identify maximum (worst-case) levels of leaching in systems that reach equilibrium before the end of shelf-life [5]. The fundamental principle behind LSERs is that solvation interactions can be decomposed into discrete, quantitatively predictable molecular interactions, providing a robust framework that transcends simple hydrophobicity measures like octanol-water partitioning [4].
Unlike traditional log-linear models that primarily correlate with octanol-water coefficients, LSERs incorporate multiple solute descriptors that account for the diverse intermolecular interactions governing partitioning behavior. This multifaceted approach enables more reliable predictions across a broad spectrum of chemical structures, from nonpolar hydrocarbons to polar compounds with significant hydrogen-bonding capacity [6]. The validation and refinement of these models require careful consideration of experimental protocols, chemical domain applicability, and data quality, all of which represent potential sources of prediction error that researchers must recognize and mitigate.
Quantitative Comparison of Model Performance
Core LSER Model Equation and Performance
The foundational LSER model for LDPE/water partitioning, as calibrated on experimental data for 156 chemically diverse compounds, is expressed by the following equation [5] [4]:
log K~i,LDPE/W~ = -0.529 + 1.098E - 1.557S - 2.991A - 4.617B + 3.886V
In this equation, the solute descriptors represent: E (excess molar refractivity), S (dipolarity/polarizability), A (hydrogen-bond acidity), B (hydrogen-bond basicity), and V (McGowan characteristic volume) [4]. This model demonstrated exceptional accuracy and precision across the training set, with reported statistics of R² = 0.991 and RMSE = 0.264 [6] [4].
Table 1: LSER Solute Descriptors and Their Interpretation in LDPE/Water Partitioning
| Descriptor | Molecular Interaction Represented | Effect on log K~i,LDPE/W~ | Physical Interpretation |
|---|---|---|---|
| E | Excess molar refractivity | Positive coefficient (+1.098) | Larger E values increase partitioning into LDPE; related to dispersion interactions |
| S | Dipolarity/polarizability | Negative coefficient (-1.557) | Higher dipolarity decreases partitioning into nonpolar LDPE |
| A | Hydrogen-bond acidity | Negative coefficient (-2.991) | Strong hydrogen-bond donors prefer aqueous phase |
| B | Hydrogen-bond basicity | Negative coefficient (-4.617) | Strong hydrogen-bond acceptors prefer aqueous phase |
| V | McGowan characteristic volume | Positive coefficient (+3.886) | Larger molecular volume favors partitioning into LDPE |
Benchmarking Against Alternative Approaches
When benchmarked against independent validation sets, the LSER approach maintains strong predictive capability, though performance varies based on the source of solute descriptors. The table below summarizes the comparative performance of different predictive approaches for LDPE/water partition coefficients.
Table 2: Performance Comparison of LDPE/Water Partition Coefficient Prediction Methods
| Prediction Method | Dataset Size | R² | RMSE | Key Limitations |
|---|---|---|---|---|
| LSER (experimental descriptors) | Independent validation (n=52) | 0.985 | 0.352 | Requires experimental solute descriptors for all compounds |
| LSER (predicted descriptors) | Independent validation (n=52) | 0.984 | 0.511 | QSPR-predicted descriptors introduce additional error |
| Log-linear (nonpolar compounds only) | Training (n=115) | 0.985 | 0.313 | Only applicable for nonpolar compounds with low H-bonding propensity |
| Log-linear (all compounds) | Training (n=156) | 0.930 | 0.742 | Poor performance for polar compounds with H-bonding capacity |
The data reveals several critical insights. First, LSER models with experimentally-derived solute descriptors provide the most robust predictions across diverse chemical classes. Second, the use of predicted descriptors, while necessary for compounds with no experimental data, introduces additional error (RMSE increases from 0.352 to 0.511) [6]. Third, traditional log-linear models based solely on octanol-water partitioning perform adequately for nonpolar compounds but show substantially degraded performance (RMSE more than doubles) when applied to polar compounds with significant hydrogen-bonding capacity [4].
Experimental Protocols for Partition Coefficient Determination
Traditional Two-Phase Equilibrium Method
The conventional approach for determining LDPE-water partition coefficients involves establishing equilibrium between purified LDPE films and aqueous solutions containing the target analytes. The general protocol involves: (1) purifying LDPE material via solvent extraction to remove manufacturing additives and impurities that might interfere with partitioning; (2) preparing analyte solutions in appropriate aqueous buffers; (3) establishing equilibrium by incubating LDPE films in analyte solutions with continuous agitation for extended periods (often 30-60 days); (4) sampling both phases post-equilibrium; and (5) quantifying analyte concentrations in both LDPE and water phases [4].
This method faces significant challenges for highly hydrophobic organic compounds (HOCs), including exceptionally long equilibration times (up to 365 days for compounds like polybrominated diphenyl ethers), difficulties in directly measuring low aqueous phase concentrations, and potential for analyte losses to vessel surfaces [7]. Studies have demonstrated that sorption of polar compounds into pristine (non-purified) LDPE can be up to 0.3 log units lower than into purified LDPE, highlighting the importance of proper polymer preparation [4].
Innovative Three-Phase System Approach
To address limitations of traditional methods, researchers have developed a novel three-phase system incorporating surfactant micelles. This approach involves adding sufficient surfactant (Brij 30) to form a micellar pseudo-phase within the polymer/water system. The K~PE-w~ values are derived from a combination of two experimentally measured values: the micelle-water partition coefficient (K~mic-w~) and the LDPE-micelle partition coefficient (K~PE-mic~) [7].
This method offers significant advantages: it reduces equilibration time to approximately half a month, avoids analytical challenges associated with direct measurement of low aqueous concentrations, and provides more precise measurements with fewer experimental errors [7]. The reliability of this approach has been demonstrated for 120 hydrophobic organic compounds, including PAHs, PBDEs, and PCBs.
Large Volume Model for Super Hydrophobic Compounds
For "super" hydrophobic organic compounds with extremely low water solubility, researchers have developed a large volume model utilizing a ~380L stainless steel container combined with dialysis tubes to generate low but steady concentrations of target analytes [31]. This system has been validated for measuring log K~pew~ values of novel halogenated flame retardants (NHFRs) and other super HOCs, overcoming solubility limitations that plague conventional methods [31].
The large volume model revealed a curvilinear relationship between log K~pew~ and log K~ow~ for target compounds, with a turning point at log K~ow~ ≈ 8.0. This nonlinearity was attributed to the large molecular volumes (>450 ų) of NHFRs, which require high Gibbs free energy to penetrate LDPE's internal structure [31].
Experimental Workflow for LSER Validation
The following diagram illustrates the comprehensive workflow for developing and validating LSER predictions for LDPE/water partitioning, integrating both experimental and computational approaches:
Diagram 1: LSER prediction validation workflow
The Scientist's Toolkit: Essential Research Reagents and Materials
Successful experimental determination of LDPE/water partition coefficients requires careful selection of materials and methods. The following table outlines key research reagents and their specific functions in partition coefficient studies.
Table 3: Essential Research Reagents and Materials for LDPE/Water Partitioning Studies
| Reagent/Material | Specification | Function in Experiment | Considerations |
|---|---|---|---|
| LDPE Film | Purified by solvent extraction | Primary polymer phase for partitioning studies | Purification critical for accurate results; pristine LDPE shows different sorption for polar compounds |
| Aqueous Buffers | pH-adjusted as needed | Aqueous phase medium; maintains physiological or environmental relevance | Buffer selection should not complex with analytes |
| Brij 30 Surfactant | Polyoxyethylene (4) lauryl ether | Forms micellar pseudo-phase in three-phase systems | Low HLB number efficient for HOCs; concentration must exceed CMC |
| Deuterated Surrogates | e.g., naphthalene-d~8~, acenaphthene-d~10~, phenanthrene-d~10~ | Internal standards for quantification | Correct for analytical recovery and matrix effects |
| Methanol/Acetone | HPLC grade | Cosolvent in cosolvent method; extraction solvent | Extrapolation to 0% cosolvent may introduce error due to nonlinearity |
| Reference Compounds | PAHs, PCBs, halogenated flame retardants | Method validation and performance reference | Established partition coefficients available for comparison |
Chemical Domain Limitations
LSER models demonstrate varying predictive accuracy across different chemical classes. The most significant errors occur when models are applied to compounds outside the chemical space represented in their training data. For LDPE/water partitioning, this is particularly relevant for polar compounds with strong hydrogen-bonding capacity, where the log-linear model shows substantially degraded performance (RMSE = 0.742 compared to 0.313 for nonpolar compounds) [4].
Mitigation Strategy: Researchers should carefully evaluate whether target compounds fall within the model's applicability domain before relying on predictions. For polar compounds outside this domain, experimental determination using the three-phase system may be preferable [7]. When extrapolation is necessary, the use of LSER models with predicted descriptors should include appropriate uncertainty estimates reflecting the increased error (RMSE ≈ 0.511) [6].
Experimental Artifacts and Method Selection
Partition coefficient measurements are susceptible to multiple experimental artifacts, including inadequate equilibration times, solute losses to container surfaces, and analytical challenges at low aqueous concentrations. Studies have shown that equilibration times can range from weeks for moderately hydrophobic compounds to over a year for super hydrophobic compounds like PBDEs [7].
Mitigation Strategy: Method selection should be guided by compound hydrophobicity. The three-phase micelle-assisted method is optimal for compounds with moderate to high hydrophobicity, reducing equilibration time to approximately half a month while maintaining accuracy [7]. For super hydrophobic compounds (log K~ow~ > 8), the large volume model provides more reliable results by maintaining stable, low aqueous concentrations [31]. Proper LDPE purification is essential, as unpurified material can yield partition coefficients up to 0.3 log units lower for polar compounds [4].
The accuracy of LSER predictions is directly dependent on the quality of solute descriptors. While experimental descriptors provide the most reliable predictions, the necessity of using predicted descriptors for compounds with no experimental data introduces additional error [6].
Mitigation Strategy: When experimental descriptors are unavailable, researchers should use QSPR prediction tools that have been validated specifically for partition coefficient prediction. The resulting predictions should include appropriate confidence intervals reflecting the additional uncertainty. For critical applications, experimental determination of descriptors or partition coefficients may be warranted.
The validation of LSER predictions for LDPE/water partitioning requires careful attention to experimental protocols, chemical domain applicability, and error source mitigation. While LSER models provide superior performance compared to traditional log-linear approaches, particularly for polar compounds, their reliability depends on appropriate application within validated chemical spaces and the use of high-quality experimental data for calibration. The development of innovative experimental methods, including three-phase micelle-assisted systems and large volume models, has addressed many traditional limitations in partition coefficient measurement, enabling more robust model training and validation. By understanding and mitigating common prediction errors through method selection, chemical domain awareness, and data quality control, researchers can leverage LSER models as powerful tools for assessing chemical distribution in pharmaceutical, environmental, and materials science applications.
The Impact of LDPE Crystallinity and Polymer Treatment on Sorption
The sorption behavior of low-density polyethylene (LDPE) is a critical determinant in its efficacy for packaging, environmental remediation, and pharmaceutical applications. This review systematically evaluates how LDPE crystallinity and various polymer treatments impact its capacity to sorb diverse chemical contaminants. Within the context of validating Linear Solvation Energy Relationship (LSER) predictions for LDPE/water partitioning, we synthesize experimental data demonstrating that crystallinity exerts a governing influence on sorption processes, often surpassing the role of particle size. Furthermore, treatments such as UV aging and the incorporation of low-molecular-weight additives significantly alter LDPE's amorphous phase and surface chemistry, thereby modulating its interactions with organic compounds. This analysis provides researchers and drug development professionals with a structured comparison of LDPE's performance relative to other polymers, underpinned by experimental sorption data and a detailed examination of the LSER predictive framework.
Low-density polyethylene (LDPE) is a cornerstone material in global industries, with the market projected to grow from US$45.2 billion in 2025 to US$64.9 billion by 2032 [32]. Its versatility, chemical resistance, and flexibility make it indispensable for packaging, agricultural films, and healthcare applications. A critical aspect of LDPE's performance across these sectors is its sorption behavior—the capacity to absorb or adsorb substances from the environment. This property directly impacts product protection, shelf life, and environmental safety, particularly in pharmaceutical development where leachable compounds must be carefully controlled.
The sorption characteristics of LDPE are not intrinsic constants but are profoundly influenced by two interconnected factors: the material's crystallinity—the proportion of structurally ordered regions within the polymer matrix—and the polymer treatments it undergoes during manufacturing or through environmental aging. These factors dictate the accessibility and chemical nature of sorption sites, thereby controlling the diffusion and partitioning of contaminants, additives, and active ingredients.
This review is framed within a broader research initiative to validate Linear Solvation Energy Relationship (LSER) predictions for LDPE/water partitioning. LSER models provide a robust quantitative framework for predicting partition coefficients based on molecular descriptors, offering tremendous utility for preemptive risk assessment in drug development. The validated LSER model for LDPE/water partitioning takes the form:
logKi,LDPE/W = -0.529 + 1.098E - 1.557S - 2.991A - 4.617B + 3.886V [6] [10] [4]
where the system parameters reflect LDPE's specific solubility interactions. Understanding how crystallinity and treatment modulate these underlying interactions is essential for applying LSER predictions accurately across different LDPE material configurations.
The Fundamental Role of Crystallinity in LDPE Sorption
Structural Basis of Sorption in Semicrystalline LDPE
LDPE is a semicrystalline polymer, characterized by a heterogeneous structure where lamellar crystals are separated by disordered amorphous regions [33]. This morphological arrangement fundamentally governs transport properties. The crystalline regions act as impermeable barriers that impede penetrant molecules, forcing them to follow tortuous pathways through the polymer matrix. In contrast, the amorphous phase serves as the primary zone for sorption and diffusion, where the free volume between polymer chains allows for the ingress and lodging of foreign molecules [33]. The efficiency of molecular packing within these amorphous regions, along with the overall crystallinity percentage, therefore becomes a decisive factor in LDPE's barrier performance and sorption capacity.
Experimental Evidence: Crystallinity as a Governing Factor
Direct experimental evidence confirms the dominant role of crystallinity in LDPE sorption behavior. A systematic investigation into the sorption of dibutyl phthalate (DBP) on polyethylene microplastics with crystallinities ranging from 17% to 99% revealed a strong inverse correlation [34]. The partition coefficient (Kd) significantly decreased with increasing crystallinity, showing a remarkable coefficient of determination (r² = 0.98) [34]. This relationship underscores that crystallinity is a more decisive factor than particle size (across a range of 27–1000 μm) in controlling the sorption of phthalate plastic additives [34].
Table 1: Impact of Polyethylene Crystallinity on Dibutyl Phthalate Sorption
| Polyethylene Type | Crystallinity (%) | Partition Coefficient, Kd (L/kg) |
|---|---|---|
| Irregularly-shaped pure PE (IPPM) | 17 | 1974.55 |
| Black plastic film PE (BPFM) | Information Missing | 1483.85 |
| White plastic film PE (WPFM) | Information Missing | 1477.45 |
| Commercial microspheres (CM) | 99 | 509.37 |
The data in Table 1 illustrates the dramatic reduction in sorption capacity as the polymer matrix becomes more crystalline. The nearly fourfold difference in Kd between the least and most crystalline materials highlights why crystallinity must be a primary consideration when predicting LDPE's interaction with its environment.
Impact of Polymer Treatments on LDPE Structure and Sorption
Modification with Low-Molecular-Weight Additives
Intentional modification of LDPE with low-molecular-weight compounds represents a strategic approach to tuning its sorption and barrier properties. Research demonstrates that incorporating additives such as tetracosane and paraffin wax into the LDPE matrix can enhance barrier properties by increasing the packing density of the amorphous regions [33]. This approach effectively reduces the free volume available for penetrant diffusion, thereby improving oxygen barrier performance.
Table 2: Effect of Additives on LDPE Properties and Sorption Behavior
| Additive Type | Concentration (wt %) | Effect on Crystallinity | Impact on Sorption/Barrier Properties |
|---|---|---|---|
| Tetracosane | 1.2 - 9.5 | Moderate increase (40.3% to 42.3%) | Improves oxygen barrier; increases packing density in amorphous regions |
| Paraffin Wax | 1.2 - 9.1 | Moderate increase (40.2% to 44.3%) | Improves oxygen barrier; increases packing density in amorphous regions |
| Paraffin Oil | 0.5 - 8.3 | Negligible change (~39.7%) | Deteriorates oxygen barrier; plasticizing effect |
The efficacy of these additives depends critically on their chemical compatibility with the LDPE matrix. Tetracosane and paraffin wax, which share structural similarity with polyethylene, integrate effectively to enhance molecular packing. In contrast, paraffin oil acts primarily as a plasticizer, potentially increasing segmental mobility and thereby compromising barrier performance despite similar crystallinity levels [33]. This distinction highlights that the chemical nature of an additive, not merely its presence, determines its impact on sorption behavior.
UV Aging and Environmental Degradation
When LDPE is exposed to environmental stressors such as ultraviolet (UV) radiation, its sorption characteristics undergo profound transformation. Accelerated UV aging induces significant structural changes in LDPE microplastics, including the formation of new functional groups such as carbonyls (-C=O), hydroxyls (-OH), and unsaturation [35]. These chemical modifications increase surface hydrophilicity and polarity, fundamentally altering the polymer's interaction with contaminants.
The sorption capacity of UV-aged LDPE for highly soluble cationic dyes like malachite green (MG) can increase up to tenfold compared to pristine LDPE [35]. This dramatic enhancement stems from a shift in the sorption mechanism. In pristine LDPE, MG sorption occurs primarily through partitioning within the amorphous regions, following a linear isotherm pattern. In UV-aged LDPE, however, the process becomes nonlinear and is dominated by adsorption at newly created polar sites on the polymer surface [35].
The crystallinity of UV-aged LDPE follows a complex trajectory, initially increasing due to chain scission in amorphous regions that allows for crystal reorganization, then eventually decreasing at higher UV doses as the crystalline regions themselves degrade [35]. This nonlinear response demonstrates the multifaceted nature of polymer aging and its consequential impact on sorption behavior.
Comparative Sorption Performance Across Polymer Types
LDPE's sorption behavior must be contextualized within the broader landscape of polymeric materials. A comparative sorption study investigating multiple polymer types with emerging contaminants revealed a distinct hierarchy of sorption affinities [36].
Table 3: Comparative Sorption percentages of Emerging Contaminants on Different Polymer Types
| Polymer Type | Sorption Order (Highest to Lowest) | Exemplary Sorption: Bisphenol A (%) | Exemplary Sorption: Progesterone (%) |
|---|---|---|---|
| Polyamide (PA) | Highest | ~80% | >40% |
| Polypropylene (PP) | High | Information Missing | >40% |
| LDPE | Medium-High | Information Missing | >40% |
| Polyvinyl Chloride (PVC) | Medium | Information Missing | Information Missing |
| High-Density PE (HDPE) | Low | Information Missing | Information Missing |
| Polyester (PES) | Lowest | Information Missing | Information Missing |
The general sorption trend across polymers was: PA > PP > LDPE > PVC > HDPE > PES [36]. This hierarchy reflects the complex interplay between polymer hydrophobicity, crystallinity, and potential for specific interactions. Polyamide's superior sorption capacity likely stems from its ability to form hydrogen bonds with contaminants, while LDPE's intermediate position reflects its balance of moderate crystallinity and flexible amorphous regions. Notably, more nonpolar contaminants such as progesterone and pyraclostrobin showed significant interaction with all polymers, including LDPE, whereas polar herbicides like atrazine and ametryn exhibited negligible sorption [36]. This confirms that hydrophobicity is a primary driver of contaminant-polymer interactions.
When compared using LSER system parameters, LDPE shows distinct behavior from polymers like polydimethylsiloxane (PDMS), polyacrylate (PA), and polyoxymethylene (POM). The latter polymers, with their heteroatomic building blocks, exhibit stronger sorption for polar, non-hydrophobic sorbates in the logKi,LDPE/W range of 3 to 4. Above this range, all four polymers converge to roughly similar sorption behavior for highly hydrophobic compounds [6] [10].
Experimental Methodologies for Sorption Analysis
Standard Sorption Experiment Protocol
The investigation of LDPE sorption typically follows a standardized experimental approach to ensure reproducibility and comparability across studies. A representative methodology for batch sorption experiments includes the following steps [36]:
LDPE Preparation: LDPE pellets or fragments are processed into microplastics (size range 63-250 μm) through grinding and sieving. Particles are washed with distilled water, sonicated to remove surface impurities, and dried at room temperature.
Contaminant Solution Preparation: Stock solutions of target contaminants (e.g., pesticides, pharmaceuticals, plasticizers) are prepared in methanol or another appropriate solvent. Working solutions are then prepared in aqueous media at relevant environmental or experimental concentrations (e.g., 0.5–10 mg L⁻¹ for DBP [34]).
Sorption Experiment: LDPE particles are combined with contaminant solutions in glass vessels at a predetermined solid-to-liquid ratio. The vessels are sealed and agitated in a temperature-controlled shaker for a specified contact time (e.g., 24 hours [36]) to reach apparent sorption equilibrium.
Phase Separation: After equilibration, the LDPE particles are separated from the aqueous phase through centrifugation or filtration.
Analytical Quantification: The contaminant concentration in the aqueous phase is quantified using analytical techniques such as high-performance liquid chromatography (HPLC) or gas chromatography-mass spectrometry (GC-MS). The sorbed amount on LDPE is calculated by the difference between initial and final aqueous concentrations.
Data Modeling: Sorption data is fitted to isotherm models (e.g., Linear, Freundlich, Langmuir) and kinetic models (e.g., pseudo-first-order, pseudo-second-order) to quantify partition coefficients and understand sorption mechanisms.
LSER Validation Framework
The validation of LSER predictions for LDPE/water partitioning follows a rigorous statistical framework [6] [10] [4]:
- Model Calibration: An initial LSER model is calibrated using a large training set of experimentally determined partition coefficients (n=156 compounds).
- Independent Validation: Approximately 33% of the total observations (n=52) are reserved as an independent validation set.
- Performance Metrics: The model's accuracy and precision are evaluated using the coefficient of determination (R²) and root mean square error (RMSE) between predicted and experimental values.
- Descriptor Source Comparison: Validation is performed using both experimentally determined LSER solute descriptors and those predicted in silico from chemical structure to assess real-world applicability.
The following diagram illustrates the experimental and computational workflow for determining and validating LDPE sorption parameters within the LSER framework:
The Scientist's Toolkit: Essential Research Reagents and Materials
Table 4: Key Research Reagents and Materials for LDPE Sorption Studies
| Reagent/Material | Function in Research | Exemplary Application |
|---|---|---|
| Dibutyl Phthalate (DBP) | Probe molecule for sorption kinetics and isotherm studies | Investigating crystallinity effects on phthalate plasticizer sorption [34] |
| Malachite Green | Model cationic, highly soluble organic dye | Studying sorption mechanism shifts in UV-aged LDPE [35] |
| Tetracosane | Low-molecular-weight additive for amorphous phase modification | Tuning oxygen barrier properties by increasing amorphous region packing density [33] |
| Bisphenol A (BPA) | Model endocrine-disrupting compound | Comparative sorption studies across multiple polymer types [36] |
| Progesterone | Model hormone contaminant | Assessing sorption of hydrophobic emerging contaminants [36] |
| Purified LDPE Powder | Controlled sorbent material with minimal additives | LSER model calibration and fundamental sorption mechanism studies [4] |
This comparison guide systematically delineates the critical factors governing sorption behavior in low-density polyethylene, with particular emphasis on crystallinity and polymer treatment. The experimental evidence unequivocally demonstrates that crystallinity is a governing factor in LDPE sorption, with partition coefficients for dibutyl phthalate showing a strong inverse correlation (r² = 0.98) with crystallinity percentage [34]. Furthermore, intentional and environmental treatments—including the incorporation of specific additives and UV aging—significantly remodel LDPE's amorphous regions and surface chemistry, thereby altering its sorption mechanisms and capacity.
When benchmarked against other common polymers, LDPE occupies an intermediate position in the sorption hierarchy (PA > PP > LDPE > PVC > HDPE > PES) [36], with its performance being highly dependent on the hydrophobicity of the target contaminant. The validated LSER model for LDPE/water partitioning provides researchers with a powerful predictive tool, but its application must be contextualized with understanding of how material properties like crystallinity influence the underlying solubility interactions.
For drug development professionals and environmental scientists, these insights are instrumental in material selection, risk assessment, and the design of advanced polymer systems. Future research directions should focus on elucidating the quantitative relationships between specific processing parameters, the resulting nanostructure of the amorphous phase, and their collective impact on LSER system parameters to enable truly predictive design of LDPE materials with tailored sorption properties.
Within pharmaceutical development, predicting the leaching of substances from plastic containers and components into drug products is a critical safety consideration. This process is largely governed by the equilibrium partition coefficient of a compound between the plastic polymer and the aqueous medium. For low density polyethylene (LDPE), a common packaging material, the Linear Solvation Energy Relationship (LSER) model provides a robust framework for predicting these partition coefficients, offering valuable insights into the behavior of chemically diverse compounds [6] [10]. This guide objectively compares the partitioning performance of polar and non-polar compounds between LDPE and water, validating the LSER model within the context of modern pharmaceutical research. The core performance gap stems from LDPE's inherent chemical nature, which is more receptive to non-polar, hydrophobic compounds, thereby presenting distinct challenges for the sorption of polar molecules [6].
Core Concepts: Polarity and the LSER Model
Fundamental Principles of Molecular Polarity
A molecule's polarity is a fundamental property dictating its physical and chemical interactions.
- Polar Bonds and Molecules: A polar covalent bond forms between atoms with significantly different electronegativities (generally >0.4), creating an uneven electron distribution and a bond dipole moment [37] [38]. A polar molecule possesses a region of partial positive charge and a region of partial negative charge, often resulting from asymmetric geometry that prevents dipole cancellation. Water (H₂O), ammonia (NH₃), and hydrofluoric acid (HF) are classic examples [39] [37].
- Non-Polar Bonds and Molecules: A non-polar covalent bond exists between atoms with identical or very similar electronegativities (difference <0.4), leading to equal electron sharing [37] [38]. A non-polar molecule has no permanent dipole moment, either because its bonds are non-polar or because its molecular symmetry causes individual bond dipoles to cancel out. Methane (CH₄), carbon dioxide (CO₂), and boron trifluoride (BF₃) fall into this category [39] [37].
The LSER Model for LDPE/Water Partitioning
The LSER model quantitatively links a compound's partitioning behavior to its fundamental molecular properties, or solute descriptors. The specific LSER model for the LDPE/water partition coefficient (( K{i, LDPE/W} )) is [6] [10]: [ \log K{i, LDPE/W} = -0.529 + 1.098E - 1.557S - 2.991A - 4.617B + 3.886V ] The variables represent the following solute descriptors:
- ( E ): Excess molar refractivity
- ( S ): Polarity/polarizability
- ( A ): Hydrogen-bond acidity
- ( B ): Hydrogen-bond basicity
- ( V ): McGowan's characteristic volume
This equation demonstrates that partitioning into LDPE is favorably influenced by a compound's volume (V), indicating a preference for larger, more hydrophobic molecules. In contrast, polarity (S), hydrogen-bond acidity (A), and hydrogen-bond basicity (B) negatively impact partitioning, as these properties increase a compound's affinity for the aqueous phase [6] [10].
Quantitative Performance Comparison
The following tables synthesize experimental and LSER-predicted data to illustrate the performance gap between polar and non-polar compounds.
Table 1: Experimental LSER Solute Descriptors and Predicted LDPE/Water Partitioning for Representative Compounds
| Compound | E | S | A | B | V | Predicted logKi, LDPE/W | Chemical Class |
|---|---|---|---|---|---|---|---|
| n-Hexane | 0.000 | 0.000 | 0.00 | 0.00 | 0.954 | 3.18 | Non-polar hydrocarbon |
| Benzene | 0.610 | 0.52 | 0.00 | 0.14 | 0.716 | 2.01 | Aromatic (weakly polar) |
| Chloromethane | 0.320 | 0.44 | 0.08 | 0.02 | 0.416 | 0.45 | Polar, H-bond donor |
| Methanol | 0.278 | 0.44 | 0.43 | 0.47 | 0.308 | -2.87 | Polar, H-bond donor/acceptor |
| Acetone | 0.179 | 0.70 | 0.04 | 0.49 | 0.547 | -1.21 | Polar, H-bond acceptor |
| Ethanol | 0.246 | 0.42 | 0.37 | 0.48 | 0.449 | -2.02 | Polar, H-bond donor/acceptor |
Table 2: Performance Benchmarking of the LSER Model (Validation Set Results) [6] [10]
| Validation Scenario | Sample Size (n) | R² | RMSE | Interpretation |
|---|---|---|---|---|
| Experimental Solute Descriptors | 52 | 0.985 | 0.352 | High precision and accuracy for known compounds |
| Predicted Solute Descriptors (QSPR) | 52 | 0.984 | 0.511 | Robust predictive power for new, unmeasured compounds |
The data in Table 1 clearly demonstrates the performance gap. Non-polar n-Hexane has a high positive logK value, indicating strong partitioning into the LDPE phase. In contrast, polar compounds like Methanol and Ethanol, which are strong hydrogen-bond donors and acceptors, have large negative logK values, reflecting their strong preference to remain in the aqueous phase. The LSER model's robust predictive capability, as shown in Table 2, makes it an invaluable tool for anticipating these behaviors across a wide chemical space.
Experimental Protocols for Model Validation
The validation of the LSER model for LDPE/water partitioning relies on a rigorous experimental methodology to generate high-quality data.
Determination of Equilibrium Partition Coefficients
The core experimental process involves measuring the concentration of a test compound in both the LDPE polymer and water phases after equilibrium has been established [6] [10].
- Sample Preparation: LDPE film is cut into standardized specimens and cleaned to remove surface contaminants. An aqueous solution of the test compound is prepared.
- Equilibration: The LDPE specimens are immersed in the solution and agitated at a constant temperature (e.g., 37°C) for a predetermined time, verified to be sufficient for reaching equilibrium.
- Phase Separation: After equilibration, the LDPE film is removed from the aqueous solution and briefly rinsed to remove any adherent solution.
- Analytical Quantification:
- The concentration of the test compound in the aqueous phase is measured directly using a suitable technique like High-Performance Liquid Chromatography (HPLC).
- The concentration in the LDPE phase is determined by either extracting the compound from the polymer and analyzing the extract, or by using a mass-balance calculation based on the concentration change in the aqueous phase.
- Calculation: The partition coefficient is calculated as ( K{i, LDPE/W} = C{LDPE} / C{W} ), where ( C{LDPE} ) and ( C_{W} ) are the equilibrium concentrations in the LDPE and water phases, respectively.
Determination of LSER Solute Descriptors
For the model to be applied, the solute descriptors (E, S, A, B, V) for each compound must be known. These can be obtained through two primary routes [6] [10]:
- Experimental Measurement: Descriptors are determined from a series of meticulously designed laboratory experiments, such as measuring gas-liquid partition coefficients, reverse-phase HPLC retention times, and solubility. These experimental values are considered the "gold standard" and are curated in dedicated databases.
- In Silico Prediction (QSPR): For compounds lacking experimental descriptors, Quantitative Structure-Property Relationship (QSPR) prediction tools are used. These software tools calculate the necessary descriptors solely from the compound's chemical structure, greatly expanding the model's applicability domain.
Comparative Sorption Behavior of Polymers
The sorption behavior of LDPE can be contextualized by comparing its LSER system parameters to those of other polymers used in medical or pharmaceutical contexts. This comparison reveals why LDPE presents a particular challenge for polar compounds.
Table 3: Comparison of Polymer Sorption Behaviors Based on LSER System Parameters [6]
| Polymer | Key Interaction Characteristics | Performance Gap Relative to LDPE |
|---|---|---|
| Low Density Polyethylene (LDPE) | Strong lipophilicity/low polarity; high affinity for non-polar compounds via dispersion forces. | Baseline - high performance for non-polar solutes. |
| Polydimethylsiloxane (PDMS) | Similar lipophilicity to LDPE; marginally higher capacity for polar interactions. | Slightly higher sorption for polar compounds. |
| Polyacrylate (PA) | Contains polar carbonyl groups; offers significant hydrogen-bond accepting capability. | Much stronger sorption for polar, non-hydrophobic compounds. |
| Polyoxymethylene (POM) | Composed of heteroatomic building blocks (-CH₂O-); capabilities for polar interactions. | Significantly stronger sorption for hydrogen-bond acidic (H-bond donor) compounds. |
The data indicates that polymers like polyacrylate (PA) and polyoxymethylene (POM), which contain heteroatoms (e.g., oxygen) in their backbone or side chains, exhibit a much greater capacity to sorb polar molecules compared to LDPE [6]. This is because these polar polymers can engage in specific interactions, such as hydrogen bonding, with polar sorbates. For a polar compound, the partition coefficient into PA or POM could be several orders of magnitude higher than into LDPE, fundamentally altering the leaching risk profile. This highlights a critical performance gap for LDPE when in contact with formulations containing polar organic molecules.
The Scientist's Toolkit
Successful application of the LSER model and experimental validation requires a set of key reagents and materials.
Table 4: Essential Research Reagents and Materials for LDPE/Water Partitioning Studies
| Item Name | Function/Description | Critical Application Note |
|---|---|---|
| LDPE Film (Standard Grade) | The polymeric phase of interest; should be of high purity and consistent thickness/crystallinity. | Material properties (e.g., density, amorphous fraction) must be characterized as they influence partitioning [6]. |
| Chemical Solutes (Analytical Grade) | A chemically diverse set of compounds with known or determinable LSER solute descriptors. | The set should cover a wide range of polarity, size, and H-bonding capabilities to robustly test the model [6] [10]. |
| HPLC-MS System | For precise and accurate quantification of solute concentrations in the aqueous phase and in polymer extracts. | Essential for generating the high-quality experimental partition coefficient data required for model validation. |
| LSER Solute Descriptor Database | A curated database (e.g., UFZ-LSER) providing experimentally derived E, S, A, B, V values for thousands of compounds. | The gold standard for input parameters; improves prediction reliability [6]. |
| QSPR Prediction Software | Software tool for estimating LSER solute descriptors directly from a compound's molecular structure. | Crucial for making predictions for compounds not listed in experimental databases, expanding the model's scope [6] [10]. |
| Constant-Temperature Agitation Incubator | To maintain a stable temperature and ensure efficient mixing during the equilibration process. | Partitioning is temperature-dependent; consistent conditions are vital for reproducible results. |
The performance gap between polar and non-polar compounds in their partitioning from water into LDPE is significant and systematic. The inherent hydrophobicity and low polarity of LDPE create a strong thermodynamic drive for the sorption of non-polar, hydrophobic compounds, while presenting a substantial barrier to the uptake of polar, hydrogen-bonding molecules. The LSER model, with its robust predictive accuracy (R² > 0.98) and strong mechanistic foundation, provides an indispensable tool for quantifying this gap [6] [10]. For researchers in drug development, this model enables the proactive identification and risk assessment of potential leachables, guiding the critical selection of container-closure systems and ensuring that the challenges posed by both polar and non-polar compounds are effectively managed for patient safety and product stability.
Recalibration Strategies and Robust Regression Methods for Improved Accuracy
Accurately predicting the partition coefficients of compounds between low-density polyethylene (LDPE) and water is crucial for environmental monitoring, pharmaceutical safety, and chemical risk assessment. Linear Solvation Energy Relationships (LSERs), also known as Polyparameter Linear Free Energy Relationships (PP-LFERs), serve as the primary predictive models for these partition coefficients [40] [5]. The reliability of these models is heavily dependent on the quality of the experimental data used for their calibration and the statistical methods employed. Recent research has highlighted inconsistencies in previously established models due to the inclusion of unreliable data, necessitating a thorough re-examination of recalibration strategies and robust regression techniques [40]. This guide compares contemporary approaches for developing robust LSER models, providing researchers with a framework for validating predictions within LDPE/water partitioning research.
Comparative Analysis of Recalibration Approaches
Foundational Principles of LSERs for Partitioning
LSERs model partition coefficients as a function of complementary solute descriptors that quantify molecular interactions. The most common form for LDPE-water systems is the EV-type equation [40]:
log K = eE + sS + aA + bB + vV + c
Here, K is the LDPE-water partition coefficient (K~LDPE/w~). The capital letters represent Abraham's solute descriptors: E (excess molar refraction), S (dipolarity/polarizability), A (hydrogen-bond acidity), B (hydrogen-bond basicity), and V (McGowan molar volume) [40] [5]. The lower-case letters (e, s, a, b, v, c) are system-specific parameters calibrated through regression analysis, reflecting the interaction properties of the LDPE phase.
Key Recalibration Strategies
Recent studies have converged on several critical strategies to improve the accuracy and reliability of LSERs.
- Rigorous Data Curation: A primary advancement involves the careful vetting of experimental partition coefficients and solute descriptors included in the calibration set. Earlier models suffered from unexpected system parameters and predictive performance because they incorporated unreliable data, with some partition coefficients deviating by more than 1 log unit from true values [40]. Modern recalibration efforts use carefully collected data from the literature, prioritizing reliability and low redundancy across a diverse set of polar and nonpolar compounds [40] [5].
- Expansion of Chemical Space: To ensure model generality, the chemical diversity of the calibration set is crucial. Successful recalibrations now utilize over one hundred compounds with varying molecular structures, molecular weights (32 to 722 g/mol), and a wide range of hydrophobicity (log K~LDPE/w~ from -3.35 to 8.36) [5] [4]. This spans over 10 orders of magnitude, significantly improving predictive performance for a broad spectrum of chemicals [40].
- Robust Regression Methodology: The application of robust regression techniques is essential for minimizing the influence of potential outliers and ensuring stable parameter estimation. This approach leads to lower root-mean-square errors (RMSE) and more mechanistically consistent system parameters [40].
The following workflow diagram illustrates the integrated process of data curation, model calibration, and validation for developing robust LSERs.
Figure 1: Workflow for Robust LSER Model Development
Quantitative Comparison of Model Performance
The effectiveness of these recalibration strategies is demonstrated by the enhanced performance of recently published models. The table below summarizes key performance metrics and system parameters from two prominent studies.
Table 1: Performance Metrics of Recalibrated LSER Models for LDPE-Water Partitioning
| Study & Model Type | Compounds (n) | R² | RMSE (log units) | Key System Parameters |
|---|---|---|---|---|
| Recalibrated PP-LFER (EV-type) [40] | >100 | N/R | 0.15 - 0.25 | Improved consistency with n-alkane system parameters. |
| LSER (Egert et al.) [5] [4] | 156 | 0.991 | 0.264 | e=1.098, s=-1.557, a=-2.991, b=-4.617, v=3.886, c=-0.529 |
N/R: Not explicitly reported in the provided search results.
The model by Egert et al. demonstrates exceptional accuracy and precision, with a low RMSE of 0.264 log units across a extensive dataset [5] [4]. The system parameters from the recalibrated PP-LFER show greater mechanistic consistency with n-alkane-water systems, as theoretically expected for the non-polar LDPE polymer [40].
Comparison with Alternative Predictive Methods
While LSERs are the most mechanistically insightful and accurate models, other methods are used for predicting LDPE-water partition coefficients.
Table 2: Comparison of LDPE-Water Partition Coefficient Prediction Methods
| Method | Principle | Advantages | Limitations | Best Use Case |
|---|---|---|---|---|
| LSER/PP-LFER | Correlates partitioning with multiple solute descriptors for different molecular interactions [40]. | High accuracy, mechanistic clarity, applicable to a wide range of polar and non-polar compounds [40] [5]. | Requires reliable solute descriptors for the target compound. | Gold standard for accurate, mechanistically sound predictions. |
| Log-Linear Model (e.g., vs. log K~O/W~) | Simple linear correlation with octanol-water partition coefficient [5]. | Simple, requires only log K~O/W~. | Poor accuracy for polar compounds; weak correlation (R²=0.930, RMSE=0.742) when polar compounds are included [5]. | Rough estimation for non-polar compounds with low H-bonding propensity. |
| QSAR Models | Correlates partitioning with molecular structure-derived descriptors [7]. | Can fill data gaps rapidly and at low cost. | Model accuracy is highly dependent on the quality of the training data [7]. | High-throughput screening of chemicals where experimental data is scarce. |
Experimental Protocols for Method Validation
Conventional Two-Phase Partitioning
The traditional method involves placing LDPE film in an aqueous solution of the target compound and agitating until equilibrium is reached [7]. The partition coefficient is then calculated from the measured concentrations in both phases. While conceptually straightforward, this method is challenging for highly hydrophobic organic compounds (HOCs) due to exceedingly low aqueous solubility, long equilibration times (up to a year), and analytical difficulties at trace concentrations, potentially leading to significant measurement errors [40] [7].
Novel Three-Phase Partitioning System
To overcome the limitations of the conventional method, a novel three-phase system has been developed [7]. This method introduces a surfactant (e.g., Brij 30) to the LDPE-water system, creating a micellar pseudo-phase.
- Procedure:
- The LDPE-micelle partition coefficient (K~PE-mic~) is measured directly. This is more accurate as concentrations in both organic phases are high.
- The micelle-water partition coefficient (K~mic-w~) is separately determined, which is also experimentally straightforward.
- The LDPE-water partition coefficient (K~PE-w~) is calculated from the two measured values: K~PE-w~ = K~PE-mic~ × K~mic-w~ [7].
- Advantages: This method sharply decreases equilibration time (to approximately half a month), avoids direct measurement of low aqueous concentrations, and yields results with little experimental error [7].
The following diagram contrasts the experimental workflows of these two key protocols.
Figure 2: Comparison of Experimental Partition Coefficient Protocols
The Scientist's Toolkit: Essential Research Reagents and Materials
Table 3: Key Materials for LDPE-Water Partitioning Experiments
| Item | Function/Description | Application Note |
|---|---|---|
| Purified LDPE | The passive sampler phase; purification via solvent extraction is critical to remove additives that can interfere with sorption [5]. | Sorption of polar compounds can be up to 0.3 log units lower in non-purified LDPE [5]. |
| Surfactant (Brij 30) | Polyoxyethylene (4) lauryl ether; used to form micelles in the novel three-phase system [7]. | Chosen for low hydrophile-lipophile balance number and efficient solubilization of HOCs [7]. |
| Performance Reference Compounds (PRCs) | Deuterated or otherwise labeled compounds added to the sampler before deployment [7]. | Used to correct for non-equilibrium conditions during field sampling and determine sampling rates. |
| Abraham's Solute Descriptors | A set of empirically derived parameters (E, S, A, B, V) that characterize a compound's interaction properties [40]. | The accuracy of LSER predictions depends on the reliability of these descriptors for the target solute [40]. |
The recalibration of LSER models for LDPE-water partitioning using robust regression methods and carefully vetted experimental data marks a significant advancement in prediction accuracy. The resulting models, such as the one by Egert et al., demonstrate high precision (RMSE ~0.26 log units) across an exceptionally wide chemical space. For experimental validation, the novel three-phase method provides a more reliable and efficient protocol for determining partition coefficients, especially for highly hydrophobic compounds. While simpler log-linear models can suffice for initial estimates of non-polar compounds, the recalibrated LSERs represent the most robust and mechanistically insightful tool for researchers and regulators requiring high-confidence predictions of chemical partitioning behavior.
In pharmaceutical development, accurately predicting the partitioning of chemicals between plastic materials and aqueous solutions is critical for assessing the risk of leachables from container-closure systems and medical devices into drug products. Low-density polyethylene (LDPE) is one of the most common polymeric materials used in pharmaceutical packaging. Traditional methods for estimating partition coefficients between LDPE and water (Ki,LDPE/W) often relied on coarse estimations or log-linear relationships with octanol-water partition coefficients (Ki,O/W), which demonstrated limited accuracy, particularly for polar compounds [5] [4].
Linear Solvation Energy Relationships (LSERs) have emerged as a high-performing predictive tool that overcomes these limitations. The foundational LSER model for LDPE/water partitioning, as established by Egert et al., is expressed by the following equation [5] [4]:
logKi,LDPE/W = -0.529 + 1.098E - 1.557S - 2.991A - 4.617B + 3.886V
This model successfully predicts partitioning behavior for a chemically diverse set of 159 compounds (Molecular Weight: 32 to 722) covering a wide range of hydrophobicity (logKi,O/W: -0.72 to 8.61) and LDPE/water partitioning (logKi,LDPE/W: -3.35 to 8.36) [5]. The model's robustness stems from its ability to account for multiple molecular interaction parameters, making it particularly valuable for novel chemicals where experimental data is lacking.
Performance Benchmarking: LSER vs. Alternative Models
Comparative Model Performance
To objectively evaluate the predictive performance of the LSER approach, we compared it against traditional log-linear models based on octanol-water partitioning. The results, summarized in Table 1, demonstrate the superior performance of the LSER methodology across diverse chemical classes.
Table 1: Performance Comparison of Predictive Models for LDPE/Water Partitioning
| Model Type | Chemical Domain | Number of Compounds (n) | R² | RMSE | Key Limitations |
|---|---|---|---|---|---|
| LSER Model [5] | Broad chemical diversity | 156 | 0.991 | 0.264 | Requires solute descriptors |
| Log-Linear Model (logK_i,O/W) [5] | Nonpolar compounds only | 115 | 0.985 | 0.313 | Limited to nonpolar chemicals |
| Log-Linear Model (logK_i,O/W) [5] | Includes polar compounds | 156 | 0.930 | 0.742 | Poor performance for polar compounds |
| LSER with Predicted Descriptors [10] | Independent validation set | 52 | 0.984 | 0.511 | Slightly higher error with QSPR-predicted descriptors |
Domain of Applicability Assessment
The benchmarking analysis reveals critical insights into model selection strategies. The traditional log-linear model based on octanol-water partitioning performs adequately for nonpolar compounds with low hydrogen-bonding propensity but shows significantly degraded performance when applied to mono-/bipolar compounds [5]. This limitation is particularly problematic for pharmaceutical applications where polar leachables are common.
The LSER model maintains consistent accuracy across both polar and nonpolar chemical domains, with the slight increase in RMSE (0.264 to 0.511) observed when using predicted instead of experimental solute descriptors still representing excellent predictive capability for novel chemicals [10]. This demonstrates the model's robustness for application to compounds without experimentally determined descriptors.
Experimental Protocols for LSER Model Development
Core Methodological Framework
The development of a robust LSER model for LDPE/water partitioning requires careful experimental design and execution. The following protocol outlines the key steps established in the referenced studies:
Polymer Preparation: LDPE material should be purified by solvent extraction to remove oligomers and additives that might interfere with partitioning measurements. Successive washing with acetonitrile, dichloromethane, and n-hexane (2 × 30 minutes each) is recommended, followed by air-drying in a clean environment [41].
Partition Coefficient Determination: Experimental determination of logK_i,LDPE/W values should be performed using a standardized approach across a chemically diverse training set. The dataset should span a wide range of molecular weights (32-722 g/mol), vapor pressures, aqueous solubilities, and polarities to ensure broad applicability [5] [4].
LSER Calibration: Using the experimental partition coefficients, the LSER model coefficients (constants and coefficients for E, S, A, B, and V descriptors) are determined through multiple linear regression analysis. The training set should contain sufficient chemical diversity to reliably determine all system parameters [5].
Model Validation: An independent validation set (approximately 33% of total observations) should be used to evaluate model predictability without overfitting. Both experimental and predicted solute descriptors should be tested to assess real-world applicability [10].
Advanced Experimental Considerations
Recent research has highlighted several factors that require special attention during experimental protocol design:
Polymer Source Variability: Partition coefficients for polyethylene from different sources can differ by up to 30%, potentially necessitating standardization of the polymer source for highly accurate predictions [41].
Cosolvency Effects: For applications involving water-ethanol mixtures (common in pharmaceutical simulating solvents), LSER-based cosolvency models have demonstrated superior performance over traditional log-linear models for predicting partition coefficients in these binary systems [14].
Amorphous Phase Correction: When comparing polymer partitioning to liquid phases, conversion of partition coefficients to logK_i,LDPEamorph/W by considering only the amorphous fraction of the polymer as the effective phase volume improves comparability to hydrocarbon/water partitioning systems [10].
The following diagram illustrates the comprehensive workflow for developing and validating a robust LSER model:
The Scientist's Toolkit: Essential Research Reagents and Materials
Successful implementation of LSER models for novel chemicals requires specific materials and computational resources. Table 2 details the essential components of the research toolkit for this field.
Table 2: Essential Research Reagents and Materials for LSER Applications
| Item/Category | Specification/Recommended Source | Function/Purpose |
|---|---|---|
| LDPE Material | Purified by solvent extraction; standardized source recommended [41] | Primary polymer phase for partitioning studies |
| Chemical Standards | 159+ compounds spanning diverse chemical classes (MW: 32-722) [5] | Training set for model development and validation |
| LSER Solute Descriptors | Experimental determination or prediction via QSPR tools [10] | Molecular parameters for LSER calculations |
| Solvent Systems | Aqueous buffers; water-ethanol mixtures for cosolvency studies [14] | Media for partitioning experiments |
| Abraham LSER Database | Freely accessible web-based curated database [10] | Source of solute descriptors and system parameters |
| QSPR Prediction Tools | Publicly available platforms (e.g., ADME@NCATS) [42] | Prediction of LSER descriptors for novel chemicals |
Molecular Insights: LSER System Parameters and Polymer Comparison
The LSER system parameters derived for LDPE/water partitioning provide valuable insights into the fundamental interactions governing solute transfer. The calibrated model: logKi,LDPE/W = -0.529 + 1.098E - 1.557S - 2.991A - 4.617B + 3.886V reveals that LDPE exhibits strong cavity formation (positive v-coefficient of 3.886) and modest lone-pair electron interactions (positive e-coefficient of 1.098) [5] [4].
Conversely, the strongly negative a- and b-coefficients (-2.991 and -4.617, respectively) indicate that LDPE is a poor solvent for hydrogen-bonding compounds compared to water. This explains why traditional log-linear models based solely on octanol-water partitioning fail for polar compounds - they cannot adequately capture these specific anti-hydrogen bonding effects [5].
When compared to other common polymers used in passive sampling and pharmaceutical applications, LDPE's unique interaction profile becomes apparent:
- Polydimethylsiloxane (PDMS): Shows similar cavity formation but different polar interaction capabilities
- Polyacrylate (PA): Exhibits stronger sorption for polar compounds due to heteroatomic building blocks
- Polyoxymethylene (POM): Offers enhanced capabilities for polar interactions compared to LDPE
For nonpolar compounds (logK_i,LDPE/W > 3-4), all four polymers exhibit roughly similar sorption behavior, but significant differences emerge for more polar substances [10]. The following diagram illustrates the key molecular interactions captured by the LSER model:
The robust performance of LSER models for predicting LDPE/water partitioning coefficients, even for novel chemicals without experimental descriptors, represents a significant advancement over traditional approaches. The high predictability (R² = 0.984, RMSE = 0.511) achieved with QSPR-predicted descriptors enables researchers to confidently apply these models to new chemical entities during early development stages [10].
For optimal implementation in pharmaceutical development, we recommend:
- Establishing standardized LDPE materials to minimize interlaboratory variability [41]
- Utilizing the freely available Abraham LSER database for descriptor acquisition [10]
- Applying LSER-based cosolvency models for water-ethanol simulating solvents [14]
- Considering polymer-specific interaction profiles when extrapolating to other polymeric materials
The expanded application domain of LSER models, supported by their robust predictive capability for novel chemicals, provides drug development professionals with a powerful tool for more accurate safety assessments of leachables, ultimately enhancing patient safety and regulatory compliance.
Benchmarking LSER Predictions Against Experiments and Alternative Models
Within pharmaceutical development and environmental science, accurately predicting the distribution of a compound between a polymer and an aqueous phase is crucial. For low-density polyethylene (LDPE), a common material in packaging and passive sampling devices, the partition coefficient (Kpew) is a key parameter. Linear Solvation Energy Relationships (LSERs) provide a powerful in-silico tool for this purpose, but their predictive power is contingent on robust experimental validation. This guide compares the performance of established and emerging methods for measuring LDPE/water partition coefficients, framing the analysis within the broader thesis that rigorous experimental benchmarking is indispensable for validating and enhancing LSER models.
Experimental Methodologies for Determining LDPE/Water Partitioning
Several experimental approaches have been developed to measure log Kpew, each with specific strengths and applications, particularly for challenging compounds like super-hydrophobic organic chemicals (HOCs).
The Large Volume Model
The Large Volume Model was developed to overcome the difficulties in measuring log Kpew for super-HOCs, such as Novel Halogenated Flame Retardants (NHFRs), which have extremely low aqueous solubility [31] [43].
- Methodology: This system uses a large stainless-steel container (e.g., ~380 L) combined with dialysis tubes to generate low, steady concentrations of target analytes. The LDPE film is exposed in this volume, and concentrations in the polymer and water are measured at equilibrium [31].
- Key Applications: It is considered robust for determining the Kpew values of super-HOCs, facilitating the application of aquatic passive sampling techniques [31] [43].
The Co-Solvent Model
The Co-Solvent Model offers an alternative method for estimating log Kpew, especially useful for very hydrophobic compounds [31].
- Methodology: This method involves estimating the Kpew by extrapolating from measured LDPE-mixture partition coefficients in solvent-water mixtures with different solvent fractions. The log Kpew is obtained by extrapolating the data to zero co-solvent concentration [31].
- Performance Note: One study found that log Kpew values for NHFRs obtained from the co-solvent model (6.51–8.89) were significantly higher than those from the large volume model, suggesting potential solvent swelling effects that may require further investigation [31].
Direct Experimental Measurement for LSER Development
The development of accurate LSER models relies on high-quality, chemically diverse experimental data.
- Methodology: An experimental dataset of partition coefficients for a wide set of chemically diverse compounds between LDPE and water is used to regress a LSER model. For example, one model was based on 156 experimental observations, with a portion of the data (e.g., 33%, n=52) reserved for independent validation [6] [44].
- Benchmarking: The experimentally derived LSER model is then benchmarked against independent data sets. Validation can be performed using either experimental LSER solute descriptors for the test compounds or descriptors predicted in-silico from the compound's structure, with the latter typically resulting in a slightly higher error [6].
Table 1: Comparison of Experimental Methods for Determining log Kpew
| Method | Principle | Key Applications | Reported Performance |
|---|---|---|---|
| Large Volume Model [31] [43] | Equilibrium partitioning in a large volume of water to maintain steady, low analyte concentrations. | Super-Hydrophobic Organic Chemicals (HOCs) like NHFRs. | Log Kpew for NHFRs: 6.27–7.34; Curvilinear relationship with log Kow observed [31]. |
| Co-Solvent Model [31] | Extrapolation from partitioning data in solvent-water mixtures to zero co-solvent. | HOCs where direct measurement is difficult. | Log Kpew for NHFRs: 6.51–8.89; Potential solvent swelling effects [31]. |
| Direct Measurement for LSER [6] | Direct measurement of partitioning for a diverse chemical set to regress a model. | Creating and validating predictive LSER models. | High precision (n=156, R²=0.991, RMSE=0.264) [6]. |
Performance Benchmarking of Predictive Models
The ultimate test for a predictive model is its performance against a rigorously measured experimental dataset.
The LSER Model for LDPE/Water Partitioning
A key LSER model for LDPE/water partitioning, derived from experimental data, demonstrates high accuracy [6] [44]:
log Ki,LDPE/W = −0.529 + 1.098E − 1.557S − 2.991A − 4.617B + 3.886V
- Performance with Experimental Descriptors: When validated against an independent set of 52 compounds using experimental LSER solute descriptors, the model maintained high performance (R² = 0.985, RMSE = 0.352) [6] [44].
- Performance with Predicted Descriptors: In a more practical but challenging scenario, using LSER solute descriptors predicted from chemical structure via a QSPR tool resulted in a slight decrease in performance (R² = 0.984, RMSE = 0.511). This statistic is considered representative for predicting partition coefficients for compounds without experimentally determined descriptors [6].
Comparison with Other Polymers
LSER system parameters allow for the comparison of LDPE's sorption behavior with other polymers like polydimethylsiloxane (PDMS), polyacrylate (PA), and polyoxymethylene (POM) [6].
- Sorption of Polar Compounds: Polymers like PA and POM, which contain heteroatomic building blocks, exhibit stronger sorption than LDPE for more polar, non-hydrophobic sorbates up to a log Ki,LDPE/W range of 3 to 4 [6].
- Sorption of Hydrophobic Compounds: Above this range, all four polymers exhibit a roughly similar sorption behavior, dominated by hydrophobic interactions [6].
Advancements with Machine Learning
Traditional LSER models can face limitations in complex scenarios. Emerging research shows that Machine Learning (ML)-assisted LSER models can significantly enhance prediction accuracy [45].
- Case Study in Adsorption: In predicting the adsorption of polyfluoroalkyl substances (PFAS) by activated carbons in complex water matrices, traditional LSER models performed poorly (R² < 0.1). In contrast, ML-assisted LSER models dramatically improved prediction accuracy (R² = 0.13–0.80). Further application of principal component regression (PCR) resulted in even more robust and accurate predictions (R² = 0.65–0.99) [45]. This demonstrates the potential of hybrid approaches for complex partitioning problems.
Table 2: Performance Benchmarking of the LSER Model and Alternatives
| Model / Approach | Validation Method | Key Performance Metrics | Implications for Predictability |
|---|---|---|---|
| LSER (Exp. Descriptors) [6] | Independent validation set (n=52) with experimental solute descriptors. | R² = 0.985, RMSE = 0.352 | High predictability for chemicals with known descriptors. |
| LSER (Pred. Descriptors) [6] | Independent validation set (n=52) with QSPR-predicted solute descriptors. | R² = 0.984, RMSE = 0.511 | Robust but less accurate prediction for new chemicals; defines expected error. |
| ML-Assisted LSER [45] | Investigation for PFAS adsorption on activated carbon. | R² = 0.65–0.99 (with PCR) vs. R² < 0.1 (Traditional LSER) | Potential for major accuracy gains in complex systems. |
The Scientist's Toolkit: Essential Research Reagents and Materials
The following reagents and materials are fundamental to conducting research in LDPE/water partitioning and LSER model development.
Table 3: Essential Research Reagents and Materials
| Item | Function / Application | Reference / Example |
|---|---|---|
| LDPE Film | The polymer sorbent phase used in passive sampling experiments and partition coefficient determination. | [31] |
| Novel Halogenated Flame Retardants (NHFRs) | Representative super-hydrophobic organic chemicals (super-HOCs) used to test and validate experimental methods. | [31] [43] |
| Polycyclic Aromatic Hydrocarbons (PAHs) & Polychlorinated Biphenyls (PCBs) | Well-studied hydrophobic reference compounds for method validation and calibration. | [31] |
| Deuterated Surrogate Standards (e.g., PAH-d) | Internal standards used for quantification and quality control in analytical chemistry. | [31] |
| LSER Solute Descriptors (E, S, A, B, V) | The core molecular parameters used to build and apply LSER models. Can be experimental or predicted. | [6] |
Visualizing the Experimental Validation Workflow
The process of developing and validating an LSER model for LDPE/water partitioning involves a structured workflow from data generation to model application, as shown in the following diagram.
Diagram 1: LSER Model Workflow
The integration of machine learning with traditional LSER approaches represents a promising frontier for improving predictive accuracy, as visualized below.
Diagram 2: ML-Enhanced LSER Approach
This comparison guide underscores a critical theme in the validation of LSER predictions for LDPE/water partitioning: the choice of experimental method directly impacts the quality of the benchmark data, which in turn defines the reliability and application domain of the predictive model. The Large Volume and Co-Solvent models provide robust pathways for generating essential data for super-HOCs, while traditional LSERs offer excellent predictability for compounds with known descriptors. The future of accurate partitioning prediction lies in the strategic integration of these rigorous experimental methods with advanced computational approaches like machine learning, creating hybrid models capable of navigating the complexity of real-world systems.
In the field of pharmaceutical development, accurately predicting the partitioning of chemicals between plastic materials and aqueous solutions is critical for assessing patient exposure to leachables. The validation of predictive models, such as Linear Solvation Energy Relationships (LSERs), relies heavily on the use of robust performance metrics. For researchers and drug development professionals, understanding the distinct roles of R-squared (R²) and Root Mean Square Error (RMSE) is essential for evaluating model quality and making reliable risk assessments. This guide provides an objective comparison of these metrics within the context of validating LSER predictions for low-density polyethylene (LDPE)/water partition coefficients, supporting robust chemical safety decisions.
Quantitative Comparison of Performance Metrics
The following table summarizes the core characteristics of R² and RMSE, two fundamental metrics used in regression analysis.
Table 1: Core Characteristics of R² and RMSE
| Metric | Definition | Interpretation | Value Range | Primary Use Case |
|---|---|---|---|---|
| R² (R-Squared) | The proportion of variance in the dependent variable that is explained by the model's independent variables [46]. | How well the predictor variables explain the variation in the response variable. A value close to 1 is desirable [47]. | 0 to 1 (or 0% to 100%); can be negative for poor models [48]. | Explaining the percentage of variance in the response variable [46]. |
| RMSE (Root Mean Square Error) | The average magnitude of the difference between predicted and observed values, in the original units of the data [49]. | The typical distance between the model's predictions and the actual observations. A value closer to 0 indicates better performance [46]. | 0 to +∞ (Highly dependent on the scale of the dependent variable) [48]. | Quantifying the average error of predictions in absolute terms [46]. |
The table below illustrates how these metrics are applied in practice, using data from a seminal study on LSER model performance in predicting LDPE/water partition coefficients.
Table 2: Performance Metrics from LSER Model Validation Studies
| Model Context | Dataset Size (n) | R² | RMSE | Interpretation |
|---|---|---|---|---|
| LSER Model Calibration (Training) [4] [5] | 156 | 0.991 | 0.264 | The model exhibits an excellent fit, explaining 99.1% of the variance in the partition coefficients with a very low average error. |
| Independent Model Validation [6] | 52 | 0.985 | 0.352 | The model maintains strong predictive power on unseen data, with a slight, expected increase in error. |
| Validation with Predicted Descriptors [6] | 52 | 0.984 | 0.511 | Using predicted instead of experimental inputs slightly increases error, but model performance remains high. |
| Log-Linear Model (Polar Compounds) [4] | 156 | 0.930 | 0.742 | This simpler model is substantially less accurate and explanatory for polar compounds, highlighting the superiority of the LSER approach. |
Interpreting Metrics in Tandem for LSER Model Validation
While Table 1 defines the metrics, their true power emerges when they are interpreted together. In the context of LSER model validation, each metric provides a different, essential piece of the performance picture.
R² for Explanatory Power: The extremely high R² value (0.991) reported during LSER model calibration indicates that the solute descriptors (E, S, A, B, V) successfully explain almost all of the systematic variance in LDPE/water partitioning behavior [4] [5]. This gives researchers confidence that the model captures the fundamental physicochemical interactions governing the partitioning process.
RMSE for Predictive Accuracy: The RMSE value of 0.264 for the same model translates to a typical prediction error of about 0.26 log units in the partition coefficient [4]. Since partition coefficients can span over 10 log units, this low RMSE indicates the model's predictions are highly precise in absolute terms, which is critical for accurate patient exposure estimates [6].
The Bottom Line for Model Comparison: Although R² is a highly informative and scale-free metric [48], the root mean squared error is often considered the primary statistic for model comparison because it directly quantifies the error in the model's predictions and determines the width of confidence intervals [50]. When comparing the LSER model to a traditional log-linear model for polar compounds, the LSER's superior performance is evidenced by its simultaneously higher R² (0.991 vs. 0.930) and lower RMSE (0.264 vs. 0.742) [4].
Experimental Protocol for Metric Evaluation
The high-quality metrics cited in the tables above are the result of a rigorous experimental and validation methodology. The following diagram outlines the key steps in this process for evaluating an LSER model.
Diagram: Workflow for LSER Model Validation. This chart outlines the key experimental stages for generating and validating a model, leading to the calculation of performance metrics.
The workflow involves several critical stages:
- Data Collection & Curation: The foundation of a robust model is high-quality experimental data. For the referenced LSER model, this involved determining partition coefficients for 159 chemically diverse compounds, ensuring coverage of a wide range of molecular weight, hydrophobicity, and polarity [4] [5].
- Model Calibration: The LSER model is fitted to the training data, deriving the coefficients for the solute descriptors in the equation
logKi,LDPE/W = -0.529 + 1.098E - 1.557S - 2.991A - 4.617B + 3.886V[4]. - Validation: The model's performance is assessed on data not used during calibration. This includes internal checks and, more importantly, validation on a fully independent set of compounds (e.g., n=52) to test its real-world predictive power [6].
- Performance Metric Calculation: Finally, R² and RMSE are calculated for both the calibration and validation sets, providing a comprehensive view of the model's fit and predictive accuracy.
Essential Research Reagent Solutions
The following table details key materials and computational tools required for conducting similar LSER partitioning studies and model validations.
Table 3: Key Reagents and Tools for LSER Partitioning Research
| Item / Solution | Function / Description | Research Context |
|---|---|---|
| Purified LDPE | The polymer phase for partitioning studies. Purification via solvent extraction removes interferents, which is critical for accurate measurement, especially for polar compounds [4]. | Material for sorption studies. |
| Chemical Solute Library | A diverse set of compounds with varying molecular weight, polarity, and hydrogen-bonding capacity to ensure the model covers a broad chemical space [4]. | Model calibration and validation. |
| Abraham Solute Descriptors (E, S, A, B, V) | Quantitative parameters describing a molecule's excess molar refraction, polarity, hydrogen-bond acidity/basicity, and volume [6]. | The independent variables in the LSER model. |
| QSPR Prediction Tool | A computational tool to predict Abraham solute descriptors for compounds where experimental measurements are unavailable [6]. | Enables model application to a wider range of chemicals. |
| Statistical Software (R, Python, RegressIt) | Platforms for performing linear regression, calculating R², RMSE, and other diagnostic statistics, and comparing models [50] [51]. | Model development and evaluation. |
Selecting and interpreting the right performance metrics is fundamental to validating predictive models in pharmaceutical development. For LSER models predicting LDPE/water partitioning, R² and RMSE are complementary metrics that together provide a complete picture of model performance. R² confirms the model captures the underlying physicochemical principles, while RMSE quantifies its predictive precision in practical, absolute terms. The benchmark values from published studies—such as R² > 0.98 and RMSE < 0.35 for independent validation—provide a high standard for researchers to aim for when developing and selecting models for critical chemical safety risk assessments.
Predicting the partitioning behavior of chemicals is a fundamental challenge in pharmaceutical and environmental sciences. For researchers investigating the migration of substances from plastic materials, accurately forecasting how compounds distribute between low-density polyethylene (LDPE) and aqueous media is particularly crucial. Two principal computational approaches dominate this predictive landscape: Linear Solvation Energy Relationships (LSERs) and Log-Linear Octanol-Water Models. This guide provides an objective comparison of these methodologies, framing the analysis within the broader thesis of validating LSER predictions specifically for LDPE/water partitioning research. We present experimental data, detailed protocols, and analytical tools to help researchers select the most appropriate model for their specific applications in drug development and material science.
Theoretical Foundations and Comparative Mechanics
Fundamental Principles of Each Approach
LSERs (Polyparameter Linear Free Energy Relationships) employ a multifaceted system that describes partitioning behavior through a set of solute descriptors representing specific molecular interactions. The general LSER form for a partition coefficient (log K) is expressed as:
log K = c + eE + sS + aA + bB + vV
Here, the capital letters represent solute-specific descriptors (E for electron interactions, S for polarity/dipolarity, A for hydrogen-bond acidity, B for hydrogen-bond basicity, V for molecular volume), while the lowercase coefficients are system-specific parameters that reflect the complementary properties of the partitioning phases [52]. For LDPE/water systems, the specific LSER model reported in recent literature is:
log Ki,LDPE/W = -0.529 + 1.098E - 1.557S - 2.991A - 4.617B + 3.886V [6] [10]
This comprehensive approach successfully captures various intermolecular interactions including dispersion, polarity/polarizability, and hydrogen bonding, offering a mechanistically grounded prediction framework.
In contrast, Log-Linear Octanol-Water Models (1p-LFERs) utilize a simplified single-parameter approach based on the hypothesis that partitioning in diverse systems correlates with the well-established n-octanol/water partition coefficient (log Kow). The general form is expressed as:
log K = m × log Kow + c
Where m and c are system-specific fitted parameters. This method essentially assumes that n-octanol serves as an adequate surrogate for complex biological and polymeric phases, with hydrophobicity as the primary driving force for partitioning [52] [53].
Visualizing the Conceptual Differences
The fundamental difference between these approaches lies in their dimensionality and the nature of molecular interactions they capture. The following diagram illustrates this conceptual distinction:
Performance Comparison and Experimental Validation
Quantitative Performance Metrics for LDPE/Water Partitioning
Direct comparative studies between LSER and log-linear models specifically for LDPE/water partitioning reveal significant differences in predictive performance. The following table summarizes key quantitative metrics from recent rigorous evaluations:
Table 1: Performance Comparison for LDPE/Water Partition Coefficient Prediction
| Model Type | Specific Model | Dataset Size (n) | R² | RMSE (log units) | Application Domain | Source |
|---|---|---|---|---|---|---|
| LSER | Experimental descriptors | 156 | 0.991 | 0.264 | Diverse organic compounds | [6] |
| LSER | Validation set | 52 | 0.985 | 0.352 | Chemically diverse validation set | [6] [10] |
| LSER | QSPR-predicted descriptors | 52 | 0.984 | 0.511 | Compounds without experimental descriptors | [6] [10] |
| Log-Linear | Not specified | 305 | 0.95* | 0.61* | Storage lipid-water partitioning | [53] |
| 2p-LFER | log Kow + log Kaw | 305 | 0.971 | 0.375 | Storage lipid-water partitioning | [53] |
Note: Values marked with * are from analogous lipid-water partitioning systems as direct LDPE/water comparisons for log-linear models were not available in the searched literature.
The performance advantage of LSER models is particularly evident for LDPE/water systems, where the complex interaction landscape of the polymeric phase requires capturing multiple interaction types. The exceptional performance (R² = 0.991, RMSE = 0.264) demonstrated by the LSER approach for LDPE/water partitioning underscores its mechanistic superiority for this specific application [6].
Chemical Domain Applicability and Limitations
The chemical space applicability of both models varies significantly, particularly for specialized compound classes:
Table 2: Chemical Domain Applicability Assessment
| Compound Class | LSER Performance | Log-Linear Performance | Key Findings | Source |
|---|---|---|---|---|
| Non-ionic surfactants | Not specifically tested | Moderate (with structural corrections) | Slow-stirring method recommended; QSPR predictions show positive bias | [54] |
| Ionizable surfactants | Not specifically tested | Poor to moderate | Significant challenges for anionic, cationic, amphoteric surfactants | [54] |
| Sulfonamides | Not specifically tested | Good (MLR models: R²=0.84, RMSE=0.64) | Multiple Linear Regression effective for this drug class | [55] |
| Diverse organic chemicals | Excellent (156 compounds) | Not specifically tested | High precision across chemically diverse set | [6] |
| Per- and polyfluoroalkyl substances | Good (via 2p-LFER) | Poor to moderate | 2p-LFER performance comparable to pp-LFER, surpassing 1p-LFER | [52] |
For ionizable compounds, the limitations of log-linear models become particularly pronounced. A recent study on surfactants concluded that "generally, QSPR-predicted log Kow/D values do not correlate well with experimental values, apart for the group of non-ionic surfactants" [54]. This highlights a fundamental limitation of single-parameter approaches for compounds exhibiting complex phase behavior or ionization.
Experimental Protocols and Methodological Considerations
Experimental Determination of Partition Coefficients
The gold-standard experimental methods for determining partition coefficients provide the crucial validation data for computational models. For LDPE/water systems specifically, the following experimental approaches are most relevant:
Slow-Stirring Method for Polymer-Water Partitioning
- Principle: Direct measurement of equilibrium distribution between LDPE and water phases under controlled, minimal-turbulence conditions [6]
- Procedure: LDPE material is immersed in aqueous solution containing test compound; system is stirred slowly (typically 150 rpm) to minimize emulsion formation; concentration in both phases measured after equilibrium establishment (typically 48-168 hours) [54]
- Phase volume ratios: Multiple ratios (e.g., 0.5:1, 1:1, 2:1 water:polymer) tested to verify equilibrium independence
- Analytical techniques: HPLC-MS, GC-MS, or direct spectroscopy for concentration quantification
- Key advantage: Minimizes microemulsion formation, particularly important for surface-active compounds [54]
- Critical consideration: Must operate below critical micelle concentration for surfactants [54]
Chromatographic Methods for log Kow Determination
- Principle: Reverse-phase HPLC retention times correlated with known log Kow values of reference compounds [56]
- Procedure: C18 or similar reverse-phase column with methanol-water or acetonitrile-water mobile phase; logarithmic retention factor (log k) extrapolated to 100% aqueous mobile phase (log kw); calibration curve established using reference compounds with known log Kow [56]
- Key advantage: High-throughput capability; minimal compound requirement
- Critical limitation: Requires appropriate reference compounds and validation for specific chemical classes [56]
Experimental Workflow Visualization
The following diagram illustrates the typical experimental and computational workflow for developing and validating partition coefficient models:
Key Research Reagent Solutions
Table 3: Essential Materials and Computational Tools for Partition Coefficient Research
| Resource Category | Specific Tool/Reagent | Function/Purpose | Application Context | Key Considerations |
|---|---|---|---|---|
| Polymer Phases | Low-Density Polyethylene (LDPE) | Representative polymeric phase for partitioning studies | Medical device leaching, packaging materials | Amorphous fraction considered as effective phase volume [6] |
| Reference Partitioning Systems | n-Octanol/Water System | Gold-standard hydrophobicity reference | Log-linear model calibration | Slow-stirring method preferred over shake-flask [54] |
| Chromatographic Materials | Silica-based C18 Columns | Stationary phase for log Kow determination | High-throughput screening | Limited stability at extreme pH (>10) [56] |
| LSER Descriptor Sources | UFZ-LSER Database | Curated source of experimental solute descriptors | LSER model development | Contains ~8,000 chemicals with experimental descriptors [52] |
| Computational Tools | EPI Suite (U.S. EPA) | log Kow and log Kaw prediction | Log-linear model inputs | Integrated QSAR models for property estimation [53] |
| Alternative Models | 2p-LFER Approaches | Balance of simplicity and accuracy | When full LSER descriptors unavailable | Uses log Kow + log Kaw [52] [53] |
Emerging Methodologies and Advanced Approaches
Deep Learning Techniques are increasingly applied to log Kow prediction, with recent models achieving RMSE values of 0.47 log units on test datasets and 0.33 for external validation sets like the SAMPL6 challenge [57]. These approaches leverage large curated datasets and can automatically extract relevant molecular features without predefined descriptors.
Hybrid 2p-LFER Models represent a promising middle ground, using linear combinations of log Kow and air-water partition coefficients (log Kaw) to capture both hydrophobicity and volatility traits of chemicals. These models have demonstrated performance comparable to full LSER approaches for protein-water partitioning (R² = 0.878, RMSE = 0.334) while maintaining simpler parameter requirements [52].
Within the specific context of validating LSER predictions for LDPE/water partitioning research, the comparative analysis strongly supports LSERs as the superior approach for this application. The demonstrated performance (R² = 0.991, RMSE = 0.264) on chemically diverse compound sets [6] underscores their mechanistic advantage for capturing the complex interactions between solutes and polymeric phases.
For research and drug development professionals, the following specific recommendations emerge:
For LDPE/water partitioning studies, LSER models should be the preferred choice when experimental solute descriptors are available or can be reliably predicted.
For high-throughput screening applications where computational efficiency is prioritized, log-linear models provide reasonable estimates within their chemical domain applicability, particularly for neutral compounds with close structural analogs in training sets.
For ionizable compounds or surfactants, exercise caution with log-linear models and consider experimental validation using slow-stirring methods.
When full LSER descriptors are unavailable, emerging 2p-LFER approaches using log Kow and log Kaw offer a balanced compromise between simplicity and accuracy.
The ongoing expansion of curated descriptor databases and development of hybrid approaches continues to enhance the accessibility and accuracy of partition coefficient prediction, providing researchers with an increasingly sophisticated toolkit for pharmaceutical and environmental applications.
Validating Linear Solvation Energy Relationship (LSER) predictions is a critical step in refining models for environmental fate and transport, particularly for LDPE/water partitioning research which informs passive sampling strategies. This process requires robust experimental benchmarking against polymer alternatives with distinct chemical characteristics. Polydimethylsiloxane (PDMS), polyacrylate, and polyoxymethylene (POM) represent widely used polymers with fundamentally different properties that influence their partitioning behavior. This guide objectively compares these polymers through synthesized experimental data, detailed methodologies, and analytical workflows to support the evaluation and refinement of LSER models for hydrophobic organic compound (HOC) partitioning.
Polydimethylsiloxane (PDMS) is an elastomeric silicon-based polymer characterized by a flexible siloxane backbone and hydrophobic methyl groups. It exhibits exceptional chain flexibility, with a glass transition temperature (Tg) typically below -120°C, which contributes to its rubbery state at ambient conditions and high permeability to gases and vapors [58]. Its inherent hydrophobicity (contact angle ~108°) and organic compound affinity make it valuable in passive sampling and microfluidic applications [58].
Polyacrylate represents a class of polymers derived from acrylate esters, with properties highly tunable based on monomer selection and side-chain chemistry. Unlike PDMS, polyacrylates typically exhibit higher Tg values and can exist in glassy or rubbery states depending on their specific composition and cross-linking density [59] [60]. Their more polar backbone and functionalizability make them suitable for coating applications, including modified versions like silicone-polyacrylate hybrids used in marine environments [61] [62].
Polyoxymethylene (POM), also known as acetal, is a semi-crystalline engineering thermoplastic with a backbone of repeating -CH2-O- units. Its high crystallinity and strong intermolecular forces result in a rigid, dense material with excellent mechanical properties and chemical resistance [7]. POM's crystalline structure creates a different partitioning environment for HOCs compared to the more flexible, amorphous regions of PDMS and polyacrylates.
Comparative Property Analysis
Table 1: Fundamental Properties of Benchmarking Polymers
| Property | PDMS | Polyacrylate | POM |
|---|---|---|---|
| Primary Backbone Chemistry | Siloxane (-Si-O-) | Carbon-carbon with ester side groups | -CH2-O- |
| Glass Transition Temperature (Tg) | < -120°C [58] | Variable (-50°C to >100°C) [59] | ~ -85°C [59] |
| Young's Modulus | 360-870 kPa [58] | Medium (MPa range) | High (GPa range) |
| Hydrophobicity (Contact Angle) | ~108° [58] | Moderate to high (modifiable) [61] | Moderate |
| Key Structural Features | Flexible backbone, hydrophobic methyl groups | Tunable side chains, functionalizable | Highly crystalline, rigid chain structure |
| Crystallinity | Amorphous [59] | Typically amorphous | Semi-crystalline [59] |
| Primary Applications in Research | Passive samplers, microfluidics, biomedical devices [58] | Protective coatings, composite materials [61] [62] | Passive samplers, engineering components [7] |
Table 2: Performance in Experimental and Modeled Contexts
| Performance Metric | PDMS | Polyacrylate | POM |
|---|---|---|---|
| LDPE-Water Partition Coefficient (Log KPE-w) Reference Range | Similar log K values to LDPE for HOCs [7] | Not commonly reported for partitioning studies | Log KPE-w values determined for PCBs, PAHs, PBDEs [7] |
| Hydrophobic Organic Compound Uptake | High affinity for non-polar compounds [58] [7] | Moderate, highly dependent on composition | High affinity, used for HOC sampling [7] |
| Chemical Resistance | Good, but swells in non-polar solvents [58] | Good to excellent when cross-linked | Excellent resistance to hydrocarbons |
| Durability in Aqueous Environments | Excellent bio-inertness, long-term stability [58] | Good, with PDMS modification enhancing marine performance [61] | Good hydrolytic stability |
| Experimental Equilibration Time | Weeks to months for HOCs [7] | Not specifically documented | Several weeks for HOCs [7] |
| LSER Model Compatibility | Well-established parameters | Limited parameters available | Established parameters for HOC partitioning [7] |
Experimental Protocols for Partition Coefficient Determination
Three-Phase Partitioning System for LDPE-Water Partitioning
A novel three-phase system developed for determining low-density polyethylene (LDPE)-water partition coefficients (KPE-w) addresses challenges in traditional two-phase methods, particularly for highly hydrophobic organic compounds (HOCs) where low aqueous solubility and long equilibration times present analytical difficulties [7].
Materials and Reagents:
- Polymer Films: LDPE, PDMS, or other test polymer films of standardized thickness
- Target Analytes: Hydrophobic organic compounds (e.g., PAHs, PCBs, PBDEs)
- Surfactant Solution: Polyoxyethylene (4) lauryl ether (Brij 30) at concentrations above critical micelle concentration
- Reference Compounds: Performance reference compounds (PRCs) for equilibrium monitoring
- Solvents: High-purity solvents for extraction and analysis (e.g., hexane, acetone)
Experimental Workflow:
- Micelle-Water Partitioning (Kmic-w): Determine micelle-water partition coefficients by measuring solute solubility in surfactant solutions above critical micelle concentration using batch equilibrium techniques [7].
- Polymer-Micelle Partitioning (KPE-mic): Equilibrate polymer films with micellar solutions containing target HOCs, then analyze both phases after equilibration.
- KPE-w Calculation: Calculate polymer-water partition coefficients using the relationship: KPE-w = KPE-mic × Kmic-w [7].
- Quality Control: Include method blanks, replicates, and PRCs to monitor equilibrium status and analytical precision.
This method significantly reduces equilibration time to approximately half a month compared to traditional methods that may require up to a year for highly hydrophobic compounds, while avoiding analytical challenges associated with direct aqueous phase concentration measurements [7].
PDMS Modification and Coating Performance Protocol
The modification of polyacrylate with PDMS demonstrates an experimental approach to enhancing polymer properties for specific applications, with relevance to partitioning behavior at interfaces [61] [62].
Synthesis Methodology:
- Emulsion Polymerization: Combine acrylate monomers (methyl methacrylate, isooctyl acrylate) with mono-terminated PDMS macromers in aqueous emulsion [62].
- Copolymer Characterization: Verify successful grafting via FTIR spectroscopy (identification of Si-CH3 peaks at 1259 cm⁻¹ and Si-O-Si at 1015 cm⁻¹) and thermal analysis [61] [62].
- Coating Application: Apply copolymer emulsions to substrates, cure to form continuous films, and condition prior to testing.
- Performance Evaluation: Assess hydrophobicity through contact angle measurements, anti-icing properties through ice adhesion tests, and durability through environmental aging cycles [62].
This solvent-free approach yields materials with surface-enriched PDMS segments that maintain flexible, hydrophobic character even after extensive environmental exposure, demonstrating the potential for creating tailored polymer interfaces with predictable partitioning behavior [62].
Experimental Workflow Visualization
Research Reagent Solutions
Table 3: Essential Research Materials for Partitioning Studies
| Reagent/Material | Function in Experiment | Application Context |
|---|---|---|
| Brij 30 Surfactant | Forms micellar pseudo-phase to enhance HOC solubility and reduce equilibration time | Three-phase partitioning systems for KPE-w determination [7] |
| Performance Reference Compounds (PRCs) | Pre-loaded reference compounds to monitor equilibrium status and determine sampling rates | Passive sampler calibration for all polymer types [7] |
| Mono-terminated PDMS (M-PDMS) | Macromer for creating graft copolymers with controlled architecture | PDMS-polyacrylate hybrid synthesis for enhanced coating performance [62] |
| Low-Density Polyethylene (LDPE) Film | Reference polymer phase with well-characterized partitioning behavior | Benchmarking against PDMS, POM, and other test polymers [7] |
| Hydrophobic Organic Compound Standards | Model analytes for partitioning experiments (e.g., PAHs, PCBs, PBDEs) | Partition coefficient determination and LSER validation [7] |
Discussion and Research Implications
The comparative data presented reveals fundamental structure-property relationships that govern polymer performance in partitioning applications. PDMS's flexible siloxane backbone and low Tg contribute to its high uptake capacity and rapid equilibration for HOCs, while POM's crystalline structure creates a more constrained partitioning environment with potentially different molecular selectivity. Polyacrylates offer intermediate properties that are highly tunable through copolymer design, as demonstrated by PDMS-modified versions that maintain hydrophobicity while improving mechanical stability [61] [62].
For LSER validation in LDPE/water partitioning research, these benchmarking studies provide critical experimental data across polymer classes with distinct solvation interactions. The three-phase partitioning method addresses key methodological challenges in obtaining reliable KPE-w values, particularly for highly hydrophobic compounds where traditional methods show significant limitations [7]. Future research directions should focus on expanding LSER models to incorporate polymer-specific structural parameters, including crystallinity, Tg, and specific chemical functionalities that influence HOC partitioning.
The integration of experimental benchmarking with computational approaches, such as the polymer informatics frameworks emerging in the field [63], promises to accelerate the development of predictive models for polymer selection in environmental monitoring and pharmaceutical applications. Standardized testing protocols and comprehensive datasets across polymer classes will be essential for validating these computational tools and ensuring their utility in real-world applications.
Independent Validation Sets and the Proof of Predictive Reliability
In the field of environmental chemistry and pharmaceutical development, the ability to predict the partition coefficients of compounds between low-density polyethylene (LDPE) and water is crucial for assessing chemical fate, transport, and bioavailability. Linear Solvation Energy Relationships (LSERs) have emerged as a powerful predictive tool for this purpose. However, the true measure of any predictive model lies not in its performance on the data used to create it, but in its proven reliability on independent, unseen data. This guide examines the critical role of independent validation sets in proving the predictive reliability of LSER models for LDPE-water partitioning, providing a comparative analysis of model performance and the experimental protocols that underpin them.
The validation process separates theoretically interesting models from those with genuine practical utility. For researchers and drug development professionals, understanding the evidence behind a model's claims is essential for selecting the right tool for applications ranging from passive environmental sampling to predicting the behavior of pharmaceutical compounds. This article objectively compares the performance of different modeling approaches based on their validated outcomes.
The Role and Structure of Independent Validation Sets
In predictive modeling, an independent validation set consists of data points that are strictly withheld from the model during its development and training phases. These points are used only once, after the model is fully calibrated, to provide an unbiased assessment of its predictive capability on new compounds. This approach prevents model overfitting—where a model appears highly accurate for the data it was trained on but fails to generalize to new data—and provides a realistic estimate of how the model will perform in real-world applications.
For LSER models predicting LDPE-water partition coefficients (logK~i,LDPE/W~), the validation process involves:
- Compound Selection: A representative subset of chemically diverse compounds is withheld from the initial model calibration.
- Descriptor Availability: Validation requires experimental LSER solute descriptors for the compounds in the validation set.
- Performance Metrics: Key statistics such as R² (coefficient of determination) and RMSE (root mean square error) are calculated by comparing predicted values against experimental measurements.
The chemical diversity of the validation set is particularly crucial, as it tests the model's ability to handle the varied molecular interactions that influence partitioning behavior across different compound classes.
Comparative Performance of LSER Models
Benchmarking LSER Predictive Accuracy
The predictive reliability of LSER models for LDPE-water partitioning has been rigorously tested using independent validation sets. The table below summarizes the performance of different modeling approaches, highlighting the importance of validation methodology:
Table 1: Performance Comparison of LSER Models for Predicting LDPE-Water Partition Coefficients
| Model Type | Validation Set Size (n) | R² | RMSE | Key Characteristics |
|---|---|---|---|---|
| LSER with Experimental Solute Descriptors [6] | 52 | 0.985 | 0.352 | High predictive accuracy for compounds with experimentally determined descriptors |
| LSER with Predicted Solute Descriptors [6] | 52 | 0.984 | 0.511 | Broader applicability but slightly reduced accuracy, suitable for screening |
| Novel Three-Phase System Model [7] | 120 | N/A | Little errors | Reduced equilibration time, avoids analytical challenges of direct measurement |
| Conventional Two-Phase Method [7] | N/A | N/A | Large discrepancies | Long equilibration times, analytical challenges for hydrophobic compounds |
Interpretation of Performance Metrics
The high R² values (approximately 0.985) demonstrate strong correlation between predicted and experimental values, indicating the models explain most of the variance in the partitioning behavior. The RMSE values (0.352-0.511) suggest that LSER predictions typically fall within approximately 0.35-0.51 log units of the true value, which is considered sufficient for many environmental and pharmaceutical applications where partition coefficients can span several orders of magnitude [6].
The slight performance difference between models using experimental versus predicted solute descriptors is noteworthy. While both approaches show excellent R² values, the higher RMSE for models using predicted descriptors reflects the additional uncertainty introduced by estimating the molecular descriptors from chemical structure alone [6]. This distinction is crucial for researchers to consider when selecting an approach for their specific application.
Experimental Protocols for Model Validation
LSER Model Development and Validation Protocol
The following diagram illustrates the comprehensive workflow for developing and validating LSER models, from initial data collection through final validation:
Three-Phase Partitioning Experimental Method
A novel experimental method for determining LDPE-water partition coefficients addresses challenges associated with conventional approaches:
Table 2: Key Research Reagents and Materials for Three-Phase Partitioning Studies
| Reagent/Material | Specifications | Function in Experimental Protocol |
|---|---|---|
| Low-Density Polyethylene (LDPE) | Single-phase polymeric material | Passive sampling phase for hydrophobic organic compounds |
| Polyoxyethylene (4) lauryl ether (Brij 30) | Nonionic surfactant, >99% purity | Forms micellar pseudo-phase to enhance solute solubility |
| Hydrophobic Organic Compounds | PAHs, PCBs, PBDEs with purity >99% | Target analytes for partition coefficient determination |
| Methanol/Acetone | HPLC grade | Polar organic solvents for cosolvent method comparisons |
Detailed Experimental Workflow:
Micelle-Water Partitioning (K~mic-w~): The relationship between total chemical solubility in Brij 30 solutions and surfactant concentrations in the micellar pseudo-phase is established. The enhancement of chemical solubility above the critical micelle concentration (cmc) is linearly dependent on Brij 30 concentrations, with line slopes representing the mass of chemical solubilized per mass of surfactant [7].
LDPE-Micelle Partitioning (K~PE-mic~): LDPE strips are immersed in chemical solutions containing surfactant at concentrations above the cmc. The system is rotated to achieve equilibrium, typically within approximately half a month—significantly shorter than traditional methods requiring up to 365 days for highly hydrophobic compounds [7].
LDPE-Water Partition Coefficient Calculation (K~PE-w~): The final partition coefficient is calculated from the combination of two experimentally measured values: K~PE-w~ = K~PE-mic~ × K~mic-w~. This approach avoids direct measurement of low aqueous concentrations, which is a major challenge for highly hydrophobic compounds [7].
Implications for Research and Development
The rigorous validation of LSER models for LDPE-water partitioning has significant implications for environmental monitoring and pharmaceutical development:
Regulatory Decision-Making: Validated models provide reliable tools for predicting environmental fate of contaminants without resource-intensive laboratory measurements for every compound [6] [7].
Drug Development Applications: For pharmaceutical researchers, these models enable prediction of compound partitioning in biological systems and drug delivery devices where polymer-water interactions are relevant.
Quality Assurance in Passive Sampling: The accuracy of polymer-water partition coefficients directly impacts the calculation of aqueous phase pollutant concentrations using performance reference compounds, essential for environmental monitoring [7].
The demonstrated predictive reliability through independent validation sets allows researchers to apply these models with greater confidence across diverse chemical domains, supporting more efficient and evidence-based scientific decision-making.
Conclusion
The validation of LSER models for LDPE/water partitioning establishes them as a superior, robust, and user-friendly tool for accurately predicting the behavior of leachables. Synthesizing the key intents, the foundational principles provide a solid theoretical base, while methodological guides enable direct application in pharmaceutical risk assessment. Troubleshooting insights ensure models are used within their optimal domain, and rigorous comparative validation confirms their performance surpasses that of traditional log-linear models, especially for chemically diverse compounds. The future direction for biomedical research involves the further integration of these predictive approaches into standardized safety protocols, the expansion of models to more complex biological phases, and the continuous refinement of descriptors through QSPR advancements. This will significantly enhance the reliability of patient exposure estimations, streamline regulatory evaluations, and minimize the resources required for comprehensive chemical risk assessments of plastic materials used in medicine and pharmaceutics.
References
- 1. Linear Solvation–Energy Relationships (LSER) and ... [https://www.mdpi.com/2673-8015/3/1/7]
- 2. In Silico Package Solutions for Deriving Values ... [https://papers.ssrn.com/sol3/papers.cfm?abstract_id=4110665]
- 3. In silico package models for deriving values of solute ... [https://pubmed.ncbi.nlm.nih.gov/36625152/]
- 4. Linear solvation energy relationships (LSERs) for robust ... [https://www.sciencedirect.com/science/article/pii/S0928098722000227]
- 5. Experimental partition coefficients and model calibration [https://pubmed.ncbi.nlm.nih.gov/35150822/]
- 6. Linear solvation energy relationships (LSERs) for robust ... [https://www.sciencedirect.com/science/article/pii/S0928098722000239]
- 7. water partition coefficients for a range of hydrophobic ... [https://www.sciencedirect.com/science/article/abs/pii/S0269749121018054]
- 8. Predicting low density polyethylene-water partition ... [https://www.sciencedirect.com/science/article/abs/pii/S0378381219304352]
- 9. water partition coefficients for a range of hydrophobic ... [https://pubmed.ncbi.nlm.nih.gov/34583266/]
- 10. Linear solvation energy relationships (LSERs) for robust ... [https://pubmed.ncbi.nlm.nih.gov/35122951/]
- 11. Predicting low density polyethylene-water partition ... [https://ui.adsabs.harvard.edu/abs/2020FlPEq.50612374Z/abstract]
- 12. The chemical interpretation and practice of linear solvation ... [https://pubmed.ncbi.nlm.nih.gov/16889784/]
- 13. Recommendations for Extractables and Leachables Testing [https://www.bioprocessintl.com/single-use/recommendations-for-extractables-and-leachables-testing]
- 14. Evaluation of Two Cosolvency Models to Predict Solute ... [https://link.springer.com/article/10.1007/s11095-022-03210-4]
- 15. Navigating Regulatory Changes in E&L Testing for 2025 [https://www.leadventgrp.com/blog/navigating-regulatory-changes-in-el-testing-for-2025]
- 16. A rapid vacuum-based method for measuring equilibrium ... [https://pubmed.ncbi.nlm.nih.gov/41161175/]
- 17. Quantifying uncertainty in predicted chemical partition ratios ... [https://pubs.rsc.org/en/content/articlehtml/2025/em/d5em00357a]
- 18. Determination of sets of solute descriptors from ... [https://pubmed.ncbi.nlm.nih.gov/15214659/]
- 19. QSPRs for Predicting Equilibrium Partitioning in Solvent– ... [https://link.springer.com/article/10.1007/s10953-022-01162-2]
- 20. Application of QSPR predictions models with experimental ... [https://www.sciencedirect.com/science/article/abs/pii/S0043135421009933]
- 21. Extraction Expertise for Medical Device Chemical ... [https://www.eag.com/blog/extraction-expertise-for-medical-device-chemical-characterization/]
- 22. Extractions: The How and Why - NAMSA [https://namsa.com/resources/audio/extractions-the-how-and-why/]
- 23. Relook on the Linear Free Energy Relationships Describing ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC7931192/]
- 24. Modeling extraction of medical device polymers for ... [https://www.sciencedirect.com/science/article/abs/pii/S0273230023000739]
- 25. Predicting partitioning from low density polyethylene to ... [https://pubmed.ncbi.nlm.nih.gov/37498141/]
- 26. Prediction of adipose tissue: Plasma partition coefficients ... [https://www.sciencedirect.com/science/article/abs/pii/S0022354916307407]
- 27. Prediction of adipose tissue: plasma partition coefficients ... [https://pubmed.ncbi.nlm.nih.gov/11170034/]
- 28. The adipose tissue:blood partition coefficient (PC [https://www.sciencedirect.com/science/article/abs/pii/S0045653599002799]
- 29. Evaluation of Two Cosolvency Models to Predict Solute ... [https://pubmed.ncbi.nlm.nih.gov/35352279/]
- 30. evaluation-of-two-cosolvency-models-to-predict-solute ... [https://www.bohrium.com/paper-details/evaluation-of-two-cosolvency-models-to-predict-solute-partitioning-between-polymers-ldpe-and-water-ethanol-simulating-solvent-mixtures/812484708771299329-9534]
- 31. Determination of low-density polyethylene–water partition ... [https://www.sciencedirect.com/science/article/abs/pii/S0045653521007049]
- 32. Low Density Polyethylene Market Trends and Forecast ... [https://finance.yahoo.com/news/low-density-polyethylene-market-trends-120300646.html]
- 33. Tuning Barrier Properties of Low-Density Polyethylene [https://pmc.ncbi.nlm.nih.gov/articles/PMC11510230/]
- 34. Implications for the behavior of phthalate plastic additives [https://www.sciencedirect.com/science/article/pii/S0269749121009751]
- 35. Impact of accelerated UV aging on chemical structure and ... [https://www.sciencedirect.com/science/article/abs/pii/S2213343725013417]
- 36. Microplastics and organic contaminants: Investigation of ... [https://www.sciencedirect.com/science/article/abs/pii/S0169772225000725]
- 37. Polar vs. Non-Polar Bonds & Molecules [https://chemistrytalk.org/polar-vs-non-polar-bonds/]
- 38. 4.4: Polar and Non-polar Covalent Bonds [https://chem.libretexts.org/Courses/Eastern_Mennonite_University/EMU%3A_Chemistry_for_the_Life_Sciences_(Cessna)/4%3A_Covalent_Bonding_and_Simple_Molecular_Compounds/4.4%3A_Polar_and_Non-polar_Covalent_Bonds]
- 39. 4.12: Shapes and Properties- Polar and Nonpolar Molecules [https://chem.libretexts.org/Bookshelves/Introductory_Chemistry/Chemistry_for_Changing_Times_(Hill_and_McCreary)/04%3A_Chemical_Bonds/4.12%3A_Shapes_and_Properties-_Polar_and_Nonpolar_Molecules]
- 40. Recalibrating polyparameter linear free energy ... [https://www.sciencedirect.com/science/article/abs/pii/S0021967323002650]
- 41. Polyethylene–Water and Polydimethylsiloxane ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9325362/]
- 42. Assay Guidance Manual for Drug Discovery: Robust or Go ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9590373/]
- 43. Determination of low-density polyethylene-water partition ... [https://pubmed.ncbi.nlm.nih.gov/33794435/]
- 44. Linear solvation energy relationships (LSERs) for robust ... [https://portal.fis.tum.de/en/publications/linear-solvation-energy-relationships-lsers-for-robust-prediction-2/]
- 45. Can Machine Learning Algorithms Enhance the Prediction ... [https://asu.elsevierpure.com/en/publications/can-machine-learning-algorithms-enhance-the-prediction-accuracy-o/]
- 46. RMSE vs. R-Squared: Which Metric Should You Use? [https://www.statology.org/rmse-vs-r-squared/]
- 47. MAE, MSE, RMSE, Coefficient of Determination, Adjusted ... [https://medium.com/analytics-vidhya/mae-mse-rmse-coefficient-of-determination-adjusted-r-squared-which-metric-is-better-cd0326a5697e]
- 48. The coefficient of determination R-squared is more ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC8279135/]
- 49. What are R² and RMSE? [https://click.clarity.io/knowledge/r2-rmse]
- 50. How to compare regression models [https://people.duke.edu/~rnau/compare.htm]
- 51. Proving statistical significance for regression R² values [https://stats.stackexchange.com/questions/308818/proving-statistical-significance-for-regression-r%C2%B2-values]
- 52. Streamlining Linear Free Energy Relationships of Proteins ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11683862/]
- 53. Exploring the role of octanol-water partition coefficient and ... [https://www.nature.com/articles/s41598-022-19452-6]
- 54. A comparison of log Kow (n-octanol–water partition coefficient ... [https://enveurope.springeropen.com/articles/10.1186/s12302-018-0176-7]
- 55. Multiple linear regression models for predicting the n‑ ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC8273033/]
- 56. Prediction of the n-Octanol/Water Partition Coefficients ... [https://www.mdpi.com/1420-3049/28/5/2270]
- 57. Exploring the octanol–water partition coefficient dataset ... [https://www.nature.com/articles/s42004-021-00528-9]
- 58. Properties and Applications of PDMS for Biomedical ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC8788510/]
- 59. 30.6: Polymer Structure and Physical Properties [https://chem.libretexts.org/Bookshelves/Organic_Chemistry/Organic_Chemistry_(Morsch_et_al.)/30%3A_Synthetic_Polymers/30.06%3A_Polymer_Structure_and_Physical_Properties]
- 60. Polymer Structure [https://www.nde-ed.org/Physics/Materials/Structure/polymer.xhtml]
- 61. Utilization of Polydimethylsiloxane (PDMS) in polymer ... [https://www.sciencedirect.com/science/article/abs/pii/S0950061823000703]
- 62. Durable and organic-solvent-free anti-icing coating ... [https://www.sciencedirect.com/science/article/abs/pii/S0032386124011935]
- 63. A Polymer Informatics Training and Testing Database [https://arxiv.org/html/2503.23491v1]